## I. INTRODUCTION
Cassava belongs to the family of Euphorbiaceae and includes 98 species. The crop is native to the American continent, being distributed from the USA to Africa. The main diversity center of cassava (Brazil) possesses at least 78 species, approximately $80\%$ of the total number of species. M. esculenta is its only domesticated species (Rogers and Appan, 1973).
Cassava plays an essential part in the food security of millions of families in tropical and subtropical regions of Africa. It is one of the main sources of carbohydrates, especially in developing regions, where it is grown as subsistence crop (FAO, 2023). Cassava has a wide range of uses in the so-called '4Fs' of: (i) food for human consumption, (ii) feed for animals, (iii) fuel, which in the form of ethanol is produced from cassava, and (iv) factories, where it is used to make alcohol, citric acid, clothing, medicines, paper, and chemicals. For many years, global demand for cassava has grown strongly due to its many industrial uses and the fact that it has often been cheaper than other starchy crops. This has then lifted it to the status of being the world's $5^{\text{th}}$ most important crop, after corn, wheat, rice, and potatoes.
In 2022, 303 million tons of cassava were produced globally worldwide, grown on 23.87 million hectares, with an average yield of $11.24\mathrm{t}\mathrm{ha}^{-1}$ (FAO, 2023). In Togo, cassava used to be a crop of the poor for a long time, but of late it is becoming more of a staple crop especially in the areas of production (Sogbedji et al., 2015). Across the country, 10,297 hectares were occupied by cassava plantations, and 38,542 tons of cassava root were harvested in 2022, with an average yield of $4\mathrm{t}\mathrm{ha}^{-1}$, $75\%$ lower than the global productivity (DSID, 2022).
Cassava is a diploid (2n=36 chromosomes) and monoicous species, with predominantly allogamous fertilization, making it highly heterozygotic (Pootakhan et al. 2014) and giving it high genetic diversity, even though it propagates vegetatively (Costa et al. 2013). Cassava can adapt to different edapho climatic conditions, such as drought and low-fertility soils (Vidal et al. 2015). Because of these characteristics, cassava cultivation is attractive to farmers with limited resources in Togo.
Despite the importance of cassava as a staple crop (Sogbedji et al., 2015), its genetic diversity is poorly documented and consequently the genetic improvement of this crop is limited in Togo (Kombaté et al., 2017). The study of Kombaté et al. (2017) using ethnobotanical survey and morphological descriptors revealed the existence of high diversity. However, ethnobiological studies involving farmers' knowledge in varietal classification have shown large variations according to Agre et al.(2017). Also, there is no consistency in the naming of varieties by farmers. This results in the possibilities of duplicates and mislabelling within the local varieties collected from farmers' fields. Additionally, the number of local varieties with different features and names, most often planted together in a single field, suggest the existence of high diversity within this crop (Siqueira et al., 2009; Rabbi et al., 2015b), which is important for plant breeding and genetic resources programs. This substantial genetic variability is due to the high heterozygosity of the crop, ease of natural cross pollination, fruit dehiscence, and to the volunteer seedlings in farmers fields (Rabbi et al., 2015b; Ceballos et al., 2016). Besides, the informal plant material exchange between farmers promotes a large number of new cultivars and expand cassava genetic diversity (Peprah et al., 2020).
From a point of breeding, small-farm cultivation of cassava is of great importance to the conservation of genetic resources. Exploring the morphological diversity of a given germplasm is fundamental to guide its conservation, management and use in conventional breeding programs (Ceballos et al., 2016).
In West Africa, genetic diversity studies have been carried out for cassava germplasm management and breeding using both morphological descriptors (Adjebeng-Danquah et al., 2016; Agre et al., 2017; Kamanda et al., 2020) and molecular markers (Rabbi et al., 2014; Soro et al., 2024). In addition, multivariate analyses allows for the simultaneous integration of data for multiple traits and has been widely used to quantify the phenotypic diversity in several crops (Kamanda et al., 2020, Soro et al., 2024).
Morphological descriptors are inexpensive and easy to record for most breeders compare to molecular markers. They are strongest determinants of taxonomic classification and agronomic value of varieties (Soyode and Oyetundi, 2009; Rabbi et al., 2015b).
The objective of this study was to explore the phenotypic diversity in a core collection of cassava cultivars based on thirty two (32) morphological traits.
## II. MATERIAL AND METHODS
### a) Plant Material
A core collection (Table 1) made of: i. one hundred (100) cultivars obtained from major cassava growing areas across the country, ii. thirty five (35) improved varieties introduced from IITA cassava breeding program, iii. seven (7) cultivars sourced from the gene bank of the Laboratory of Virology and Biotechnology of the University of Lome and iv. two (2) varieties obtained from the cassava gene bank of Embrapa Mandioca Fruticultura (Cruz das Almas, BA, Brazil) was used in this study. Five improved varieties (high yielding and CMD resistant) namely Gbazekoute, TMS 96_0409, TMS 96_0166, CRI Sika Bankye and CRI Ampong bankye, recently released by the national cassava breeding unit were used as checks.
### b) Experimental Site
The experiment was run at the Togolese Agronomic Research Institute (ITRA) station of Davie (latitude: $6^{\circ}23'$ 5" North; longitude: $1^{\circ}12'$ 18" East; altitude: 76 meters) located in the cassava production belt. This site is representative of typical cassavagrowing conditions in Togo and is characterized by a bimodal rainfall pattern. During the experimentation, a total rainfall of $1231.5\mathrm{mm}$ was recorded for 80 rainy days. July was the highest monthly rainfall with 207.8 mm for 14 rainy days, while November was the lowest monthly rainfall with $8.7\mathrm{mm}$ for 4 days rainy days. The annual average temperature was $28.5^{\circ}\mathrm{C}$. The egitation is characterized by herbaceous vegetation (Banito et al., 2010). The site's soil, suitable for cassava cultivation (Ezui, 2017) and known as 'Terres de Barre,' is characterized as sandy-clay with $70\%$ sand, $3.8\%$ silt, $8.1\%$ clay, acid pH ( $\mathsf{H}_2\mathsf{O}$ 1:1) 5.5, $1.05\%$ organic matter, $0.41\%$ total nitrogen (N), 10 ppm available phosphorus (P), and cation exchange capacity (CEC) of 2.89 milliequivalents (meq)/100g of soil in the top 15 cm samples (Sogbedji et al. 2015).
### c) Experimental Design, Field Layout and Maintenance
The experiment was laid out in an augmented block design with one hundred forty four (144) cultivars as tested genotypes and five (5) checks varieties, distributed in twelve (12) blocks. Each block was delimited after ploughing and harrowing of the site. Distance of $1.5\mathrm{m}$ separated adjacent blocks and plots. The experimental unit was composed of four rows of 4 m with 16 plants of a genotype. A spacing of $1\mathrm{m}$ between plants and rows was adopted. The experiment was carried under rainfed conditions. Neither herbicide nor fertilizers were applied. The experiment was kept weed free by regular hand weeding. The trial was harvested twelve months after planting.
Table 1: List of Togo's Cassava Germplasm Cultivars Characterized
<table><tr><td>N°</td><td>Cultivar</td><td>Type</td><td>Origin</td><td>N°</td><td>Cultivar</td><td>Type</td><td>Origin</td></tr><tr><td>1</td><td>CRI Sika Bankye</td><td>Improved</td><td>Ghana</td><td>27</td><td>TMS 92_0326</td><td>Improved</td><td>Togo</td></tr><tr><td>2</td><td>CRI Ampong Bankye</td><td>Improved</td><td>Ghana</td><td>28</td><td>TMS 96_1708</td><td>Improved</td><td>Togo</td></tr><tr><td>3</td><td>TMS 95_0166</td><td>Improved</td><td>IITA</td><td>29</td><td>TMS 98_2132</td><td>Improved</td><td>Togo</td></tr><tr><td>4</td><td>TMS 96_0409</td><td>Improved</td><td>IITA</td><td>30</td><td>TMS 99_0554</td><td>Improved</td><td>Togo</td></tr><tr><td>5</td><td>Gbazekoute</td><td>Landrace</td><td>Togo</td><td>31</td><td>Agbede</td><td>Landrace</td><td>Togo</td></tr><tr><td>6</td><td>TMS 01_0006</td><td>Improved</td><td>IITA</td><td>32</td><td>Agou</td><td>Landrace</td><td>Togo</td></tr><tr><td>7</td><td>TMS 00_0354</td><td>Improved</td><td>IITA</td><td>33</td><td>Aguidagba</td><td>Landrace</td><td>Togo</td></tr><tr><td>8</td><td>TMS 00_0364</td><td>Improved</td><td>IITA</td><td>34</td><td>Akaleyo</td><td>Landrace</td><td>Togo</td></tr><tr><td>9</td><td>TMS 01_0034</td><td>Improved</td><td>IITA</td><td>35</td><td>Akebou</td><td>Landrace</td><td>Togo</td></tr><tr><td>10</td><td>TMS 01_0046</td><td>Improved</td><td>IITA</td><td>36</td><td>Akoss</td><td>Landrace</td><td>Togo</td></tr><tr><td>11</td><td>TMS 01_0093</td><td>Improved</td><td>IITA</td><td>37</td><td>Ankra atihe</td><td>Landrace</td><td>Togo</td></tr><tr><td>12</td><td>TMS 01_0098</td><td>Improved</td><td>IITA</td><td>38</td><td>Akpadjin Feto</td><td>Landrace</td><td>Togo</td></tr><tr><td>13</td><td>TMS 01_0131</td><td>Improved</td><td>IITA</td><td>39</td><td>Alagno</td><td>Landrace</td><td>Togo</td></tr><tr><td>14</td><td>TMS 01_0379</td><td>Improved</td><td>IITA</td><td>40</td><td>Ankra 3</td><td>Landrace</td><td>Togo</td></tr><tr><td>15</td><td>TMS 01_1085</td><td>Improved</td><td>IITA</td><td>41</td><td>Ankra Atiyibo</td><td>Landrace</td><td>Togo</td></tr><tr><td>16</td><td>TMS 01_1086</td><td>Improved</td><td>IITA</td><td>42</td><td>Assiatoe</td><td>Landrace</td><td>Togo</td></tr><tr><td>17</td><td>TMS 01_1097</td><td>Improved</td><td>IITA</td><td>43</td><td>Atidjin1</td><td>Landrace</td><td>Togo</td></tr><tr><td>18</td><td>TMS 01_1206</td><td>Improved</td><td>IITA</td><td>44</td><td>Atidjin 2</td><td>Landrace</td><td>Togo</td></tr><tr><td>19</td><td>TMS 01_1224</td><td>Improved</td><td>IITA</td><td>45</td><td>Atidjin Poli</td><td>Landrace</td><td>Togo</td></tr><tr><td>20</td><td>TMS 01_1368</td><td>Improved</td><td>IITA</td><td>46</td><td>Atidokpo</td><td>Landrace</td><td>Togo</td></tr><tr><td>21</td><td>TMS 01_1368(2)</td><td>Improved</td><td>IITA</td><td>47</td><td>Atihe1</td><td>Landrace</td><td>Togo</td></tr><tr><td>22</td><td>TMS 01_1371</td><td>Improved</td><td>IITA</td><td>48</td><td>Atiyibo 1</td><td>Landrace</td><td>Togo</td></tr><tr><td>23</td><td>TMS 01_1610</td><td>Improved</td><td>IITA</td><td>49</td><td>Atiyobo2</td><td>Landrace</td><td>Togo</td></tr><tr><td>24</td><td>TMS 01_1662</td><td>Improved</td><td>IITA</td><td>50</td><td>Awou</td><td>Landrace</td><td>Togo</td></tr><tr><td>25</td><td>TMS 01_1797</td><td>Improved</td><td>IITA</td><td>51</td><td>Awouye</td><td>Landrace</td><td>Togo</td></tr><tr><td>N°</td><td>Cultivar</td><td>Type</td><td>Origin</td><td>N°</td><td>Cultivar</td><td>Type</td><td>Origin</td></tr><tr><td>53</td><td>Badjogou</td><td>Landrace</td><td>Togo</td><td>79</td><td>Kanbom Bantchi</td><td>Landrace</td><td>Togo</td></tr><tr><td>54</td><td>Bazoka</td><td>Landrace</td><td>Togo</td><td>80</td><td>Kanigbeli 1</td><td>Landrace</td><td>Togo</td></tr><tr><td>55</td><td>Bob</td><td>Landrace</td><td>Togo</td><td>81</td><td>Kanigbeli 2</td><td>Landrace</td><td>Togo</td></tr><tr><td>56</td><td>Bob Assou</td><td>Landrace</td><td>Togo</td><td>82</td><td>Kataoli</td><td>Landrace</td><td>Togo</td></tr><tr><td>57</td><td>Bob Yegue</td><td>Landrace</td><td>Togo</td><td>83</td><td>Katawole</td><td>Landrace</td><td>Togo</td></tr><tr><td>58</td><td>BRS Caipira</td><td>Landrace</td><td>Brazil</td><td>84</td><td>Kidirondi</td><td>Landrace</td><td>Togo</td></tr><tr><td>59</td><td>Degaule</td><td>Landrace</td><td>Togo</td><td>85</td><td>Kisseimou Koutowou</td><td>Landrace</td><td>Togo</td></tr><tr><td>60</td><td>Djakoagni</td><td>Landrace</td><td>Togo</td><td>86</td><td>Kola</td><td>Landrace</td><td>Togo</td></tr><tr><td>61</td><td>Djeble</td><td>Landrace</td><td>Togo</td><td>87</td><td>Kolaoung</td><td>Landrace</td><td>Togo</td></tr><tr><td>62</td><td>Djolaoba</td><td>Landrace</td><td>Togo</td><td>88</td><td>Kolmon kamkam</td><td>Landrace</td><td>Togo</td></tr><tr><td>63</td><td>Djoliba</td><td>Landrace</td><td>Togo</td><td>89</td><td>Kossikouma</td><td>Landrace</td><td>Togo</td></tr><tr><td>64</td><td>Donmoyibo</td><td>Landrace</td><td>Togo</td><td>90</td><td>Koutowou 2</td><td>Landrace</td><td>Togo</td></tr><tr><td>65</td><td>Fetonegbodji</td><td>Landrace</td><td>Togo</td><td>91</td><td>Kperoung Felgou</td><td>Landrace</td><td>Togo</td></tr><tr><td>66</td><td>Flawavi</td><td>Landrace</td><td>Togo</td><td>92</td><td>Kperoung Mamogue</td><td>Landrace</td><td>Togo</td></tr><tr><td>67</td><td>Gabonvi-ESA</td><td>Landrace</td><td>Togo</td><td>93</td><td>Kpla</td><td>Landrace</td><td>Togo</td></tr><tr><td>68</td><td>Gbadovi</td><td>Landrace</td><td>Togo</td><td>94</td><td>Loki</td><td>Landrace</td><td>Togo</td></tr><tr><td>69</td><td>Gbazé- ESA</td><td>Landrace</td><td>Togo</td><td>95</td><td>M'beou</td><td>Landrace</td><td>Togo</td></tr><tr><td>70</td><td>Vivigbaze</td><td>Landrace</td><td>Togo</td><td>96</td><td>MM96/5280</td><td>Improved</td><td>Togo</td></tr><tr><td>71</td><td>Ghana spana</td><td>Landrace</td><td>Togo</td><td>97</td><td>MM96/JW2</td><td>Improved</td><td>Togo</td></tr><tr><td>72</td><td>Gnidou</td><td>Landrace</td><td>Togo</td><td>98</td><td>Nigeria Fleur</td><td>Landrace</td><td>Togo</td></tr><tr><td>73</td><td>Hogninvo 1</td><td>Landrace</td><td>Togo</td><td>99</td><td>Nigeria Kikpaou</td><td>Landrace</td><td>Togo</td></tr><tr><td>74</td><td>Hogninvo 2</td><td>Landrace</td><td>Togo</td><td>100</td><td>Nigeria Kissaimon</td><td>Landrace</td><td>Togo</td></tr><tr><td>75</td><td>Inconnu</td><td>Landrace</td><td>Togo</td><td>101</td><td>N'tossou</td><td>Landrace</td><td>Togo</td></tr><tr><td>76</td><td>IRAT- Davie</td><td>Landrace</td><td>Togo</td><td>102</td><td>Ankra atihe</td><td>Landrace</td><td>Togo</td></tr><tr><td>77</td><td>Jhonson</td><td>Landrace</td><td>Togo</td><td>103</td><td>Okpoli</td><td>Landrace</td><td>Togo</td></tr><tr><td>78</td><td>Kalba</td><td>Landrace</td><td>Togo</td><td>104</td><td>Pela</td><td>Landrace</td><td>Togo</td></tr><tr><td>No</td><td>Cultivar</td><td>Type</td><td>Origin</td><td>No</td><td>Cultivar</td><td>Type</td><td>Origin</td></tr><tr><td>105</td><td>Peloumkoute</td><td>Landrace</td><td>Togo</td><td>131</td><td>D00_126</td><td>Improved</td><td>IITA</td></tr><tr><td>106</td><td>Penivi</td><td>Landrace</td><td>Togo</td><td>132</td><td>D00_54</td><td>Improved</td><td>IITA</td></tr><tr><td>107</td><td>Sabe</td><td>Landrace</td><td>Togo</td><td>133</td><td>D00_166</td><td>Improved</td><td>IITA</td></tr><tr><td>108</td><td>Sankara</td><td>Landrace</td><td>Togo</td><td>134</td><td>Toma 9</td><td>Landrace</td><td>Togo</td></tr><tr><td>109</td><td>Sassakawa</td><td>Landrace</td><td>Togo</td><td>135</td><td>CVTM4</td><td>Landrace</td><td>Togo</td></tr><tr><td>110</td><td>Sorad</td><td>Landrace</td><td>Togo</td><td>136</td><td>Toma 162</td><td>Landrace</td><td>Togo</td></tr><tr><td>111</td><td>Sawa</td><td>Landrace</td><td>Togo</td><td>137</td><td>Unknown 02</td><td>Landrace</td><td>Togo</td></tr><tr><td>112</td><td>Spana Assou</td><td>Landrace</td><td>Togo</td><td>138</td><td>TMS 96_1317</td><td>Improved</td><td>Togo</td></tr><tr><td>113</td><td>Spana Yegue</td><td>Landrace</td><td>Togo</td><td>139</td><td>TMS 96_0304</td><td>Improved</td><td>Togo</td></tr><tr><td>114</td><td>BRS Tapioqueira</td><td>Landrace</td><td>Brazil</td><td>140</td><td>TMS 96_0102</td><td>Improved</td><td>Togo</td></tr><tr><td>115</td><td>Tassiodo</td><td>Landrace</td><td>Togo</td><td>141</td><td>TMS 96_0869</td><td>Improved</td><td>Togo</td></tr><tr><td>116</td><td>Tchigouevi</td><td>Landrace</td><td>Togo</td><td>142</td><td>TMS 96_1642</td><td>Improved</td><td>Togo</td></tr><tr><td>117</td><td>Tetetidadjin</td><td>Landrace</td><td>Togo</td><td>143</td><td>TMS 96_0590</td><td>Improved</td><td>Togo</td></tr><tr><td>118</td><td>TME 419</td><td>Improved</td><td>Togo</td><td>144</td><td>TMS 96_539</td><td>Improved</td><td>Togo</td></tr><tr><td>119</td><td>TM1</td><td>Improved</td><td>Togo</td><td>145</td><td>TMS 96_1565</td><td>Improved</td><td>Togo</td></tr><tr><td>120</td><td>TME1</td><td>Improved</td><td>Togo</td><td>146</td><td>TMS 96_0603</td><td>Improved</td><td>Togo</td></tr><tr><td>121</td><td>TME 696</td><td>Improved</td><td>Togo</td><td>147</td><td>TMS 30572</td><td>Improved</td><td>IITA</td></tr><tr><td>122</td><td>Touwevi</td><td>Landrace</td><td>Togo</td><td>148</td><td>KPEM_10_03</td><td>Improved</td><td>Togo</td></tr><tr><td>123</td><td>Tuaka Atsu</td><td>Landrace</td><td>Togo</td><td>149</td><td>TMS 4(2) 1425</td><td>Improved</td><td>IITA</td></tr><tr><td>124</td><td>Tuaka komi Mami</td><td>Landrace</td><td>Togo</td><td>129</td><td>D00_208</td><td>Improved</td><td>IITA</td></tr><tr><td>125</td><td>Yabaka</td><td>Landrace</td><td>Togo</td><td>130</td><td>D00_14</td><td>Improved</td><td>IITA</td></tr><tr><td>126</td><td>D00_8300</td><td>Improved</td><td>IITA</td><td>52</td><td>Unknown</td><td>Landrace</td><td>Togo</td></tr><tr><td>127</td><td>M94_0583</td><td>Improved</td><td>IITA</td><td></td><td></td><td></td><td></td></tr><tr><td>128</td><td>D00_137</td><td>Improved</td><td>IITA</td><td></td><td></td><td></td><td></td></tr></table>
### d) Phenotypic Data Collection
Thirty two (32) morphological traits were recorded using the cassava descriptor (Guevara et al., 2010) at three (3), six (6), nine (9) and twelve (12) months after planting (MAP). Data were recorded from the plants within the whole plot, and the most frequent occurrence variant was noted. At 12 MAP, the inner eight (8) plants within each plot were uprooted and observations on roots were taken. The traits assessment date, and method of assessment are summarized in Table 2.
Table 2: List of Morphological Traits recorded in Togo's Cassava Germplasm
<table><tr><td>N°</td><td>Trait</td><td>Code</td><td>Assessment date</td><td>1Assessment scale</td></tr><tr><td>1</td><td>Colour of apical leaves</td><td>ColApLea</td><td>3 MAP</td><td>3, 5, 7 or 9</td></tr><tr><td>2</td><td>Pubescence on apical leaves</td><td>PubApLea</td><td>3 MAP</td><td>0 or 1</td></tr><tr><td>3</td><td>Lobe margins</td><td>LoMar</td><td>6 MAP</td><td>3 or 5</td></tr><tr><td>4</td><td>Colour of leaf vein</td><td>ColLeaVe</td><td>6 MAP</td><td>3, 5, 7 or 9</td></tr><tr><td>5</td><td>Petiole Colour</td><td>PetCol</td><td>6 MAP</td><td>1, 2, 3, 5, 7 or 9</td></tr><tr><td>6</td><td>Leaf color</td><td>LeaCol</td><td>6 MAP</td><td>3, 5, 7 or 9</td></tr><tr><td>7</td><td>Number of leaf lobes</td><td>NLeaLo</td><td>6 MAP</td><td>3, 5, 7, 9 or 11</td></tr><tr><td>8</td><td>Shape of central leaflet</td><td>ShaCeLea</td><td>6 MAP</td><td>1-10</td></tr><tr><td>9</td><td>Orientation of petiole</td><td>OriPet</td><td>6 MAP</td><td>1, 3, 5 or 7</td></tr><tr><td>10</td><td>Flowering hability</td><td>FIHa</td><td>6 MAP</td><td>0 or 1</td></tr><tr><td>11</td><td>Pollen</td><td>Pol</td><td>6 MAP</td><td>0 or 1</td></tr><tr><td>12</td><td>Leaf retention</td><td>LeaRet</td><td>6 MAP</td><td>1-5</td></tr><tr><td>13</td><td>Stipule margin</td><td>StiMar</td><td>9 MAP</td><td>1 or 2</td></tr><tr><td>14</td><td>Length of stipule</td><td>LenSti</td><td>9 MAP</td><td>3 or 5</td></tr><tr><td>15</td><td>Color of stem cortex</td><td>ColStCor</td><td>9 MAP</td><td>1-3</td></tr><tr><td>16</td><td>Colour of stem epidermis</td><td>ColStEpi</td><td>9 MAP</td><td>1, 2, 3, or 4</td></tr><tr><td>17</td><td>Colour of stem exterior</td><td>ColStExt</td><td>9 MAP</td><td>3, 4, 5, 6, 7, 8 or 9</td></tr><tr><td>18</td><td>Colour of end branches of adult plant</td><td>CoEBran</td><td>9 MAP</td><td>3, 5, or 7</td></tr><tr><td>19</td><td>Growth habit of stem</td><td>GrHaSt</td><td>9 MAP</td><td>1 or 2</td></tr><tr><td>20</td><td>Distance between leaf scars</td><td>DisLeaSca</td><td>9 MAP</td><td>3, 5, or 7</td></tr><tr><td>21</td><td>Prominence of foliar scars</td><td>ProFoSca</td><td>9 MAP</td><td>3 or 5</td></tr><tr><td>22</td><td>Fruit</td><td>Frt</td><td>9 MAP</td><td>0 or 1</td></tr><tr><td>23</td><td>Levels of branching</td><td>LeBran</td><td>12 MAP</td><td>0, 1, 2 or 3</td></tr><tr><td>24</td><td>Branching habit</td><td>BranHab</td><td>12 MAP</td><td>1, 2, 3 or 4</td></tr><tr><td>25</td><td>Root constrictions</td><td>RoCons</td><td>12 MAP</td><td>1-3</td></tr><tr><td>26</td><td>Colour of root cortex</td><td>ColRoCor</td><td>12 MAP</td><td>1-4</td></tr><tr><td>27</td><td>Colour of root pulp</td><td>ColRoPu</td><td>12 MAP</td><td>1-5</td></tr><tr><td>28</td><td>External colour of storage root</td><td>ExColRo</td><td>12 MAP</td><td>1-4</td></tr><tr><td>29</td><td>Extent of root peduncle</td><td>ExRoPed</td><td>12 MAP</td><td>0, 3 or 5</td></tr><tr><td>30</td><td>Shape of plant</td><td>ShaPI</td><td>12 MAP</td><td>1-4</td></tr><tr><td>31</td><td>Root shape</td><td>RoSha</td><td>12 MAP</td><td>1-4</td></tr><tr><td>32</td><td>Texture of root epidermis</td><td>TexRoEpi</td><td>12 MAP</td><td>3, 5, or 7</td></tr></table>
### e) Phenotypic Diversity Analyses
The morphological diversity of the core collection was assessed following two approaches. Traits distribution was determined using Microsoft Excel (2016) in the first approach. In the second approach, morphological data were subjected to Multiple Correspondence Analysis (MCA) for identification of relevant traits contributing mostly to the germplasm diversity (Giles et al. 2018). From the MCA results, traits that presented the highest variability were used as active variables to perform cluster analysis for morphotypes identification within the germplasm using the Ward's method (Kawuki et al. 2011). The optimal number of clusters was determined using the distribution of the variance function methods. The morphological diversity of the germplasm was visualized by plotting the factors scores for individual genotype in the first factorial plan in order to assess the relationship among cultivars (Selamawit Abebe et al. 2021). Analyses were run in SAS version 9.4.
## III. RESULTS
### a) Descriptive Analysis of Morphological Traits
The variability observed for qualitative traits among cassava cultivars is given in figure 1. In all $37.58\%$ cultivars showed purplish green colour, $22.82\%$ had purple, $3.36\%$ showed dark green and $36.24\%$ had purplish green colour. About $45\%$ of the cultivars had pubescence on apical leaves, while $55\%$ had not (Figure 1). Approximately $25\%$ of cultivars had yellowish green petioles, $22.82\%$ purple petioles, $16.78\%$ red petioles, $15.54\%$ reddish-green petioles, $11.41\%$ green petioles and $8.72\%$ accessions showed greenish-red petioles (Figure 1). Nearly half of the cultivars $(47.65\%)$ had green leaf vein, $28.19\%$ cultivars showed reddish-green leaf vein in less than half of the lobe, $13.42\%$ had reddish-green leaf vein in more than half of the lobe and $10.74\%$ had red leaf vein. Four morphotypes were observed in the germplasm based on the leaves colour. The first morphotype with dark green leaves was represented by $51.68\%$ of the cultivars, the second morphotype had light green leaves and was represented by $22.82\%$ of the cultivars, the third morphotype exhibited purple green leaves and was represented by $14.76\%$ of the cultivars and the fourth one had purple leaves was represented by $10.74\%$ of the cultivars. The petioles of most cultivars in the collection were horizontal $(50.34\%)$, the irregular type was observed in $29.53\%$ cultuvers. $12.75\%$ cultivars showed petioles inclined downwards while $7.38\%$ cultuvers had petioles inclined upward. About $40.27\%$ accessions had lanceolate central leaflet, $32.88\%$ had elliptic-lanceolate, $10.07\%$ accessions had oblong-lanceolate central leaflet, $10.74\%$ had obovate-lanceolate, $4.03\%$ had ovoid, $0.67\%$ had linear, $0.67\%$ had pandurate and $0.67\%$ had linear pandurate central leaflet (Figure 1). Most cultivars $(66.44\%)$ showed a winding lobe margin and $33.56\%$ cultivars in the germplasm had a smooth lobe margin. In the germplasm, $61.07\%$ cultivars had nine leaf lobes, $28.19\%$ had seven leaf lobes, $8.05\%$ had eleven and $2.69\%$ five leaf lobes. For the leaves retention trait, $31.54\%$ cultivars exhibited very poor leaf retention, $33.56\%$ cultivars showed less than the average leaf retention, $19.46\%$ exhibited average leaf retention while $15.44\%$ cultivars exhibited better than average leaf retention (Figure 1).
With regards to stem related traits, $31.54\%$ cultivars showed an upright growth habit of the stems, while the cultivars exhibiting a zigzag growth habit were observed in $68.46\%$ cultivars. About $44\%$ of the characterized cultivars had green end branches, $42.95\%$ cultivars had green-purple and $12.75\%$ had purple end branches. Four morphotypes were observed in the germplasm with regard to the stem epidermis colour (Figure 1). About $33.56\%$ cultivars had light brown stem epidermis, $25.5\%$ had dark brown, $24.83\%$ had orange and $16.11\%$ showed light-green stem epidermis. The colours of stem cortex observed were dark green $(61.07\%$ cultivars), orange $(22.82\%$ cultivars) and light green $(16.11\%$ cultivars). Approximately $30\%$ cultivars had silver stem exterior, $17.45\%$ had greenish-yellowish stem exterior, $14.17\%$ showed gray stem, $12.08\%$ orange stem, $10.74\%$ dark brown stem, $9.39\%$ light brown stem and $6.04\%$ golden stem exterior. Majority of the cultivars (63.09%) exhibited prominent foliars scars while 36.91% had semi- prominent foliar scars. The distance between leaf scars was short for 92.63% cultivars characterized, medium for 6.04% cultivars and long for 1.34% cultivars. About 69.80% of the cultivars had long stipules while 30.20% cultivars had short stipules. The stipule margin of 61.74% cultivars was split or forked and entire for 38.26% cultivars (Figure 1).
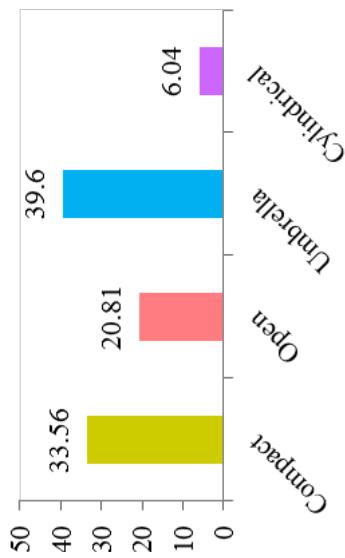
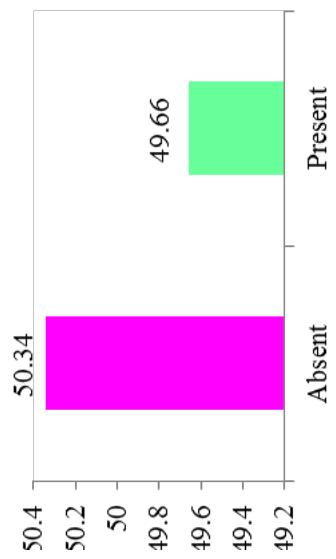
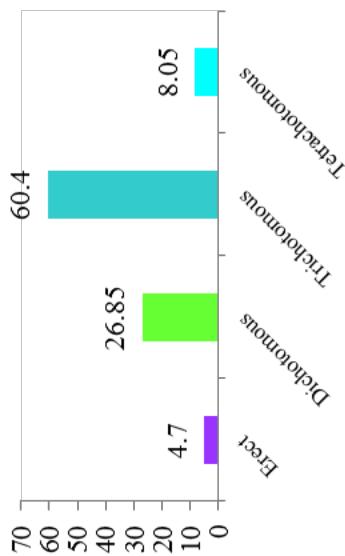
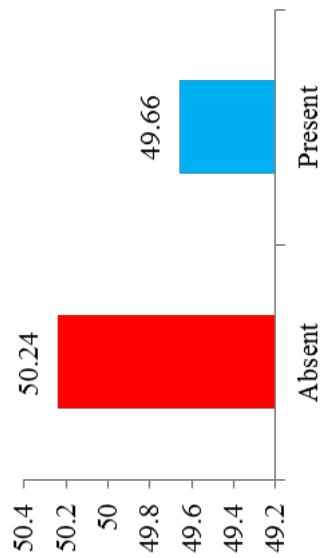
Differences in the flowering ability among cultivars were observed. About $50\%$ of the cultivars produced flowers while remaining did not flower. At harvest, seeds were observed on $42.28\%$ of cultivars. Cultivars exhibiting zero level of branching $(47\%)$ and three level of branching $(34.23\%)$ were predominant. About $60.40\%$ cultivars showed trichotomous branching habit, $26.85\%$ cultivars showed dichotomous type, 8 cultivars had tetrachotomous type while only 4 cultivars showed an erect type. Cassava cultivars examined phenotypically based on their plant shape exhibited variation with umbrella $(39.60\%$ cultivars), compact $(33.56\%)$, open $(12.64\%)$ and cylindrical $(6.04\%)$ shapes. (Figure 1)
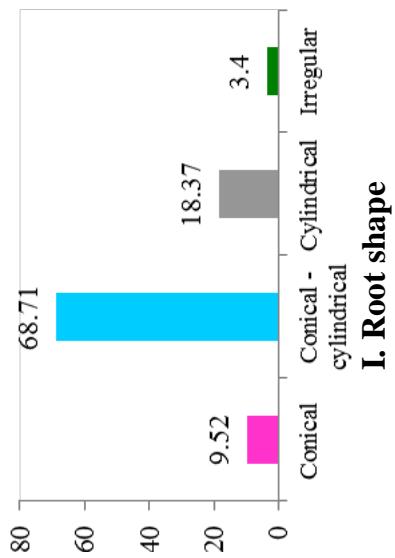
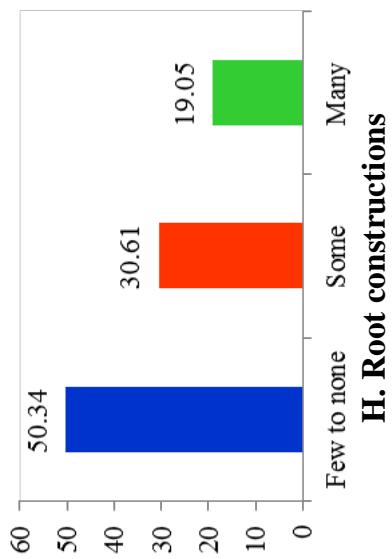
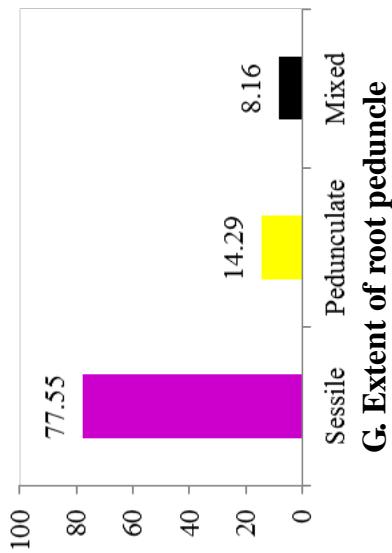
At harvest, the external colour of storage root also exhibited variation with dark brown (40.82%), light brown (35.37%), yellow (12.93%), and white or cream (10.88%) colours. The root cortex colour showed high variability among cultivars and four morphotypes were observed. Majority (49.0%) had white or cream root cortex colour, followed by yellow (22.2%), pink (11.1%) and purple (8.9%) colours. Root epidermis was white or cream for 47.62% cultivars, yellow (18%), pink (20.41%) and purple (6.52%). Based on the colour of root pulp cultivars exhibited variation with cream (48.98%), yellow (30.61%), white (14.29%), and pink (6.12%) colours. In the germplasm, 69% cultivars had conical cylindrical root shape, 18.37% cylindrical, 9.52% conical and 3.4% had irregular root shape. Cultivars with few (50.34%) and some (30.61%) root constrictions were predominant. Majority of the cultivars (77.55%) had sessile roots, whereas 14.29% had pedunculate roots. The mixed type was recorded on 8.16% of the cultivars. The texture of root epidermis exhibited variation with rough (33.33%), smooth (34%), and intermediate (32.65%) textures. (Figure 1).
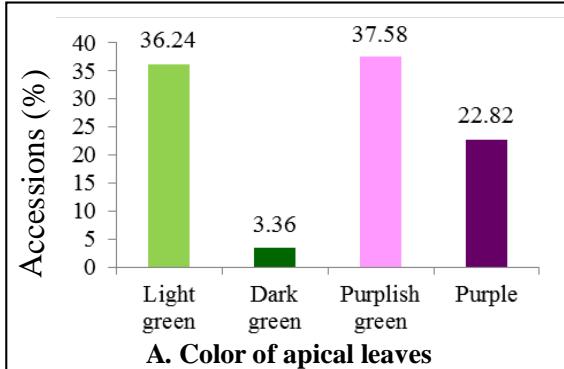
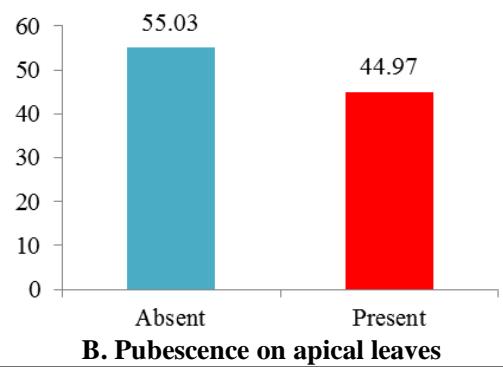 Figure 1: Frequency Distribution of 149 Cassava Cultivars based on Morphological Traits
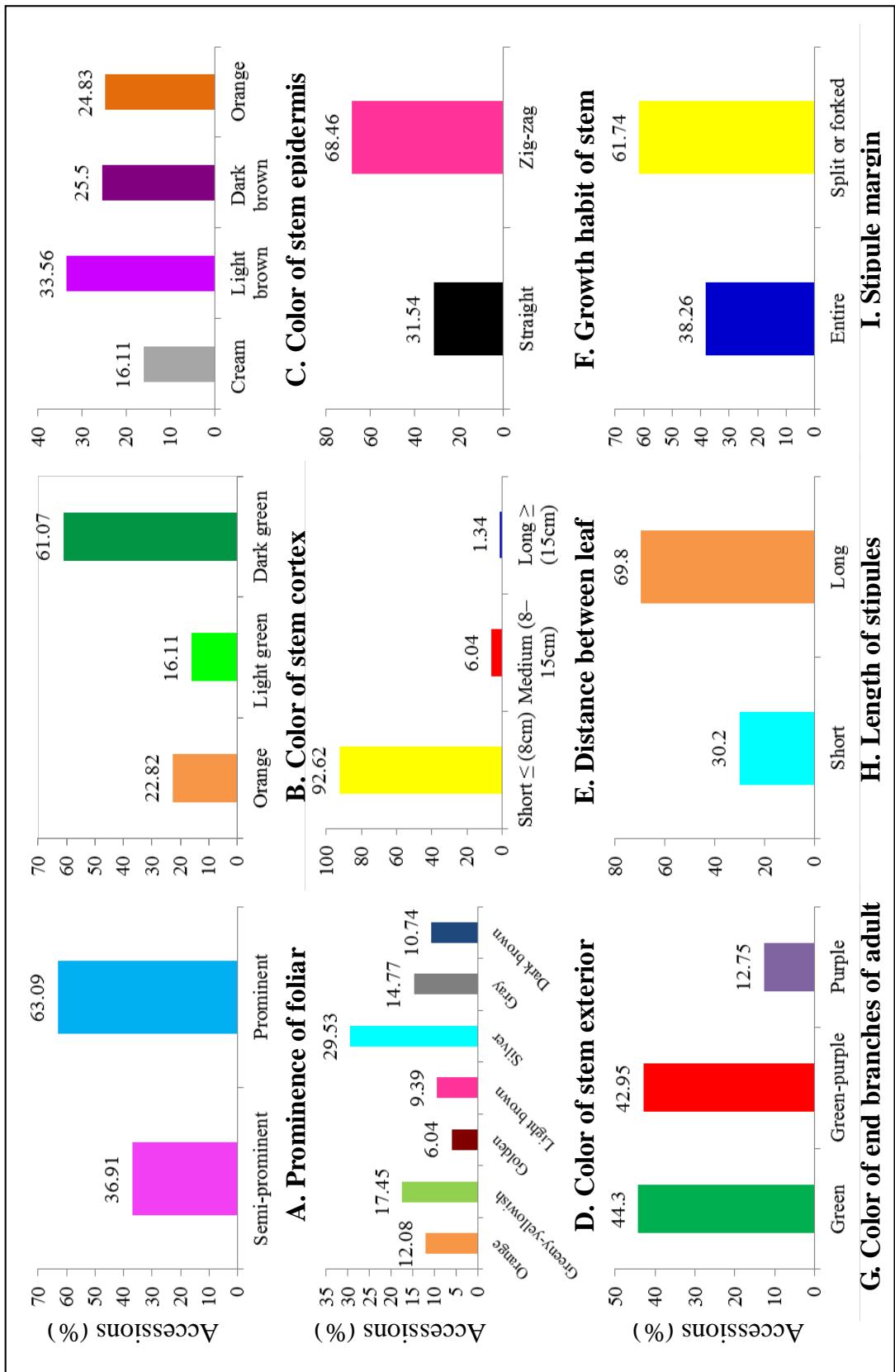 Figure 1. Count Frequency Distribution of 149 Cassava Cultivars based on Morphological Traits
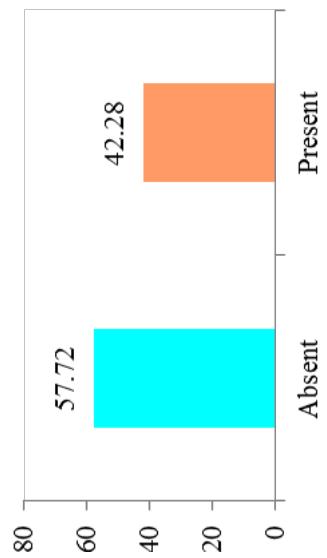 C. Fruit
 F. Shape of plant
 B. Pollen
 E. Branching habit
 eeess A. Flowering
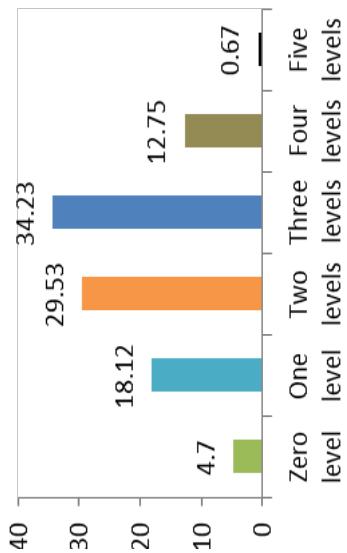 eeess D. Levels of branching
 eeess Figure 1. Frequency Distribution of 149 Cassava Cultivars based on Morphological Traits
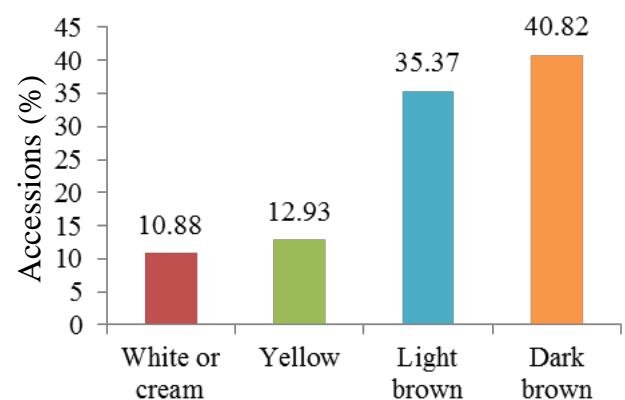 M. Exernal color of storage root
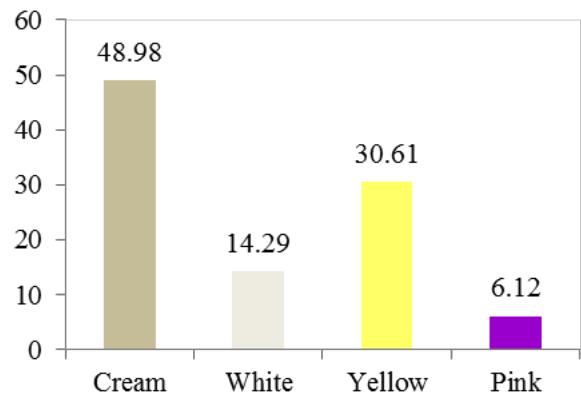 N. Color of root pulp
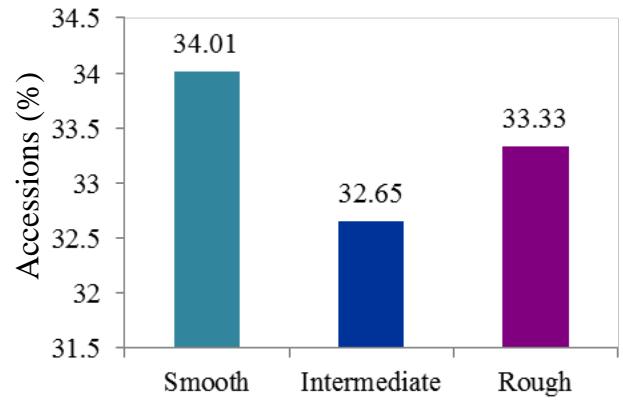 O. Color of root cortex
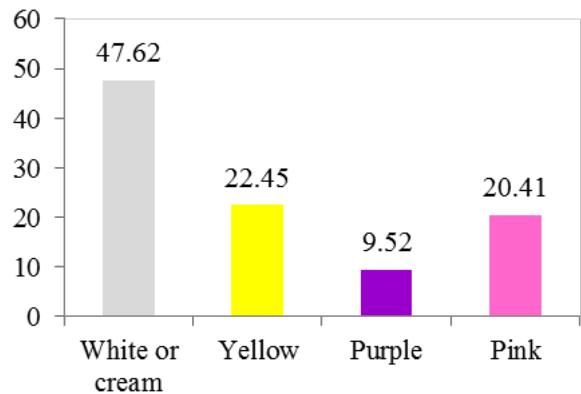 P. Color of root epidermis Figure 1: Frequency Distribution of 149 Cassava Cultivars based on Morphological Traits
### b) Diversity Among Cultivars and Differentiation based on Morphological Traits
i. Significant traits describing the germplasm diversity The objective of Multiple Correspondence Analysis (MCA) is to provide interpretable visualization of complex variable space. The meaning given to the axes and analysis of proximities between traits and conditions are usually made from the factorial planes. Thus, the first factorial plan and the factors having eigenvalue greater than one were retained. On the basis of this criterion, the first 11 factors with an eigenvalue greater than one were significant and therefore retained for the subsequent analyses (Table 3). These first eleven factors (Fs explained $68.14\%$ of the morphological variability among cultivars.
Factor 1 with an eigenvalue of 5.17 and accounted for $16.16\%$ of the morphological variability. This factor was strongly correlated with petiole colour, leaf colour, colour of leaf vein, flowering and seed set ability and the levels of branching. Factor 2 with an eigenvalue of 3, explained $9.40\%$ of the total variation, and was positively defined by leaves colour of end branches, color of stem epidermis, leaf lobe margin and the growth habit of stem. Factor 3 represented by traits such the colour of stem exterior and color of stem cortex, had an eigenvalue of 2.5, and explained $7.81\%$ of the divergence among cultivars. Factor 4 with an eigenvalue of 1.73 correlated with leaf retention and color of apical leaf. Factor 5 with an eigenvalue of 1.71 was related mainly to the distance between leaf scars. In Factor 6, shape of central leaflet and pubescence on apical leaves were the main traits, while the extent of root peduncle was most important trait in factor 7. In factor 8, the most important traits describing the germplasm variability were the stipule length and the prominence of foliar scars. Factor 9 with an eigenvalue of 1.26, contribute $3.95\%$ of the total variability and was mainly related to the stipule margin. Factor 10 was mainly represented by the orientation of petioles, plante shape and the color of root cortex while factor 11 was represented by the colour of root pulp (Table 3).
Table 3: Eigen values, proportion of variation and contribution associated with the axes of the MCA of 32 qualitative traits
<table><tr><td></td><td>F1</td><td>F2</td><td>F3</td><td>F4</td><td>F5</td><td>F6</td><td>F7</td><td>F8</td><td>F9</td><td>F10</td><td>F11</td></tr><tr><td>Eigenvalue</td><td>5.172</td><td>3.000</td><td>2.502</td><td>1.734</td><td>1.718</td><td>1.508</td><td>1.394</td><td>1.362</td><td>1.265</td><td>1.148</td><td>1.003</td></tr><tr><td>Variability (%)</td><td>16.161</td><td>9.375</td><td>7.817</td><td>5.419</td><td>5.369</td><td>4.714</td><td>4.356</td><td>4.256</td><td>3.953</td><td>3.586</td><td>3.134</td></tr><tr><td>Cumulative%</td><td>16.161</td><td>25.536</td><td>33.353</td><td>38.773</td><td>44.142</td><td>48.855</td><td>53.211</td><td>57.467</td><td>61.420</td><td>65.007</td><td>68.141</td></tr><tr><td>ColApLea1</td><td>0.053</td><td>0.122</td><td>0.001</td><td>0.287</td><td>0.006</td><td>0.010</td><td>0.016</td><td>0.040</td><td>0.037</td><td>0.051</td><td>0.068</td></tr><tr><td>PubApLea2</td><td>0.001</td><td>0.019</td><td>0.091</td><td>0.040</td><td>0.001</td><td>0.180</td><td>0.047</td><td>0.146</td><td>0.016</td><td>0.003</td><td>0.009</td></tr><tr><td>LeaRet3</td><td>0.024</td><td>0.122</td><td>0.000</td><td>0.298</td><td>0.028</td><td>0.019</td><td>0.117</td><td>0.018</td><td>0.029</td><td>0.012</td><td>0.027</td></tr><tr><td>ShaCeLea4</td><td>0.001</td><td>0.000</td><td>0.049</td><td>0.047</td><td>0.020</td><td>0.230</td><td>0.034</td><td>0.053</td><td>0.052</td><td>0.025</td><td>0.021</td></tr><tr><td>PetCol5</td><td>0.494</td><td>0.235</td><td>0.028</td><td>0.002</td><td>0.005</td><td>0.003</td><td>0.039</td><td>0.001</td><td>0.001</td><td>0.012</td><td>0.002</td></tr><tr><td>LeaCol6</td><td>0.426</td><td>0.282</td><td>0.003</td><td>0.000</td><td>0.001</td><td>0.043</td><td>0.013</td><td>0.010</td><td>0.015</td><td>0.031</td><td>0.000</td></tr><tr><td>NuLeaLob7</td><td>0.352</td><td>0.005</td><td>0.036</td><td>0.001</td><td>0.018</td><td>0.159</td><td>0.038</td><td>0.012</td><td>0.058</td><td>0.000</td><td>0.006</td></tr><tr><td>LobMar8</td><td>0.003</td><td>0.187</td><td>0.000</td><td>0.016</td><td>0.142</td><td>0.023</td><td>0.007</td><td>0.021</td><td>0.149</td><td>0.003</td><td>0.001</td></tr><tr><td>ColLeaVei9</td><td>0.409</td><td>0.352</td><td>0.002</td><td>0.000</td><td>0.015</td><td>0.027</td><td>0.012</td><td>0.008</td><td>0.017</td><td>0.016</td><td>0.008</td></tr><tr><td>OrPet10</td><td>0.014</td><td>0.142</td><td>0.061</td><td>0.023</td><td>0.001</td><td>0.027</td><td>0.146</td><td>0.008</td><td>0.000</td><td>0.215</td><td>0.013</td></tr><tr><td>Flow11</td><td>0.714</td><td>0.095</td><td>0.013</td><td>0.042</td><td>0.000</td><td>0.004</td><td>0.003</td><td>0.013</td><td>0.000</td><td>0.007</td><td>0.017</td></tr><tr><td>Pol12</td><td>0.714</td><td>0.095</td><td>0.013</td><td>0.042</td><td>0.000</td><td>0.004</td><td>0.003</td><td>0.013</td><td>0.000</td><td>0.007</td><td>0.017</td></tr><tr><td>Fru13</td><td>0.635</td><td>0.082</td><td>0.012</td><td>0.065</td><td>0.002</td><td>0.001</td><td>0.002</td><td>0.022</td><td>0.000</td><td>0.001</td><td>0.007</td></tr><tr><td>ProFoSca14</td><td>0.104</td><td>0.001</td><td>0.001</td><td>0.000</td><td>0.072</td><td>0.148</td><td>0.000</td><td>0.224</td><td>0.036</td><td>0.065</td><td>0.003</td></tr><tr><td>ColSteCor15</td><td>0.113</td><td>0.084</td><td>0.232</td><td>0.028</td><td>0.101</td><td>0.107</td><td>0.047</td><td>0.047</td><td>0.034</td><td>0.001</td><td>0.000</td></tr><tr><td>ColSteEpi16</td><td>0.014</td><td>0.187</td><td>0.018</td><td>0.021</td><td>0.086</td><td>0.173</td><td>0.001</td><td>0.005</td><td>0.009</td><td>0.057</td><td>0.013</td></tr><tr><td>ColSteExt17</td><td>0.016</td><td>0.079</td><td>0.301</td><td>0.039</td><td>0.154</td><td>0.038</td><td>0.017</td><td>0.036</td><td>0.012</td><td>0.010</td><td>0.001</td></tr></table>
Table 3: Continued: Eigen values, proportion of variation and contribution associated with the axes of the MCA of 32 qualitative traits
<table><tr><td></td><td>F1</td><td>F2</td><td>F3</td><td>F4</td><td>F5</td><td>F6</td><td>F7</td><td>F8</td><td>F9</td><td>F10</td><td>F11</td></tr><tr><td>Eigenvalue</td><td>5.172</td><td>3</td><td>2.502</td><td>1.734</td><td>1.718</td><td>1.508</td><td>1.394</td><td>1.362</td><td>1.265</td><td>1.148</td><td>1.003</td></tr><tr><td>Variability (%)</td><td>16.161</td><td>9.375</td><td>7.817</td><td>5.419</td><td>5.369</td><td>4.714</td><td>4.356</td><td>4.256</td><td>3.953</td><td>3.586</td><td>3.134</td></tr><tr><td>Cumulative%</td><td>16.161</td><td>25.536</td><td>33.353</td><td>38.773</td><td>44.142</td><td>48.855</td><td>53.211</td><td>57.467</td><td>61.420</td><td>65.007</td><td>68.141</td></tr><tr><td>\(DisLeaSca^{18} \)</td><td>0.105</td><td>0.001</td><td>0.010</td><td>0.061</td><td>0.189</td><td>0.009</td><td>0.023</td><td>0.052</td><td>0.085</td><td>0.052</td><td>0.042</td></tr><tr><td>\(GroHaSte^{19} \)</td><td>0.036</td><td>0.179</td><td>0.025</td><td>0.002</td><td>0.028</td><td>0.011</td><td>0.105</td><td>0.070</td><td>0.031</td><td>0.000</td><td>0.031</td></tr><tr><td>\(ColBrAdPl^{20} \)</td><td>0.275</td><td>0.319</td><td>0.006</td><td>0.074</td><td>0.045</td><td>0.011</td><td>0.048</td><td>0.007</td><td>0.002</td><td>0.001</td><td>0.012</td></tr><tr><td>\(LenSti^{21} \)</td><td>0.058</td><td>0.020</td><td>0.061</td><td>0.096</td><td>0.005</td><td>0.002</td><td>0.019</td><td>0.243</td><td>0.011</td><td>0.002</td><td>0.081</td></tr><tr><td>\(StiMar^{22} \)</td><td>0.007</td><td>0.000</td><td>0.021</td><td>0.031</td><td>0.018</td><td>0.005</td><td>0.110</td><td>0.001</td><td>0.489</td><td>0.055</td><td>0.016</td></tr><tr><td>\(LevBran^{23} \)</td><td>0.211</td><td>0.193</td><td>0.022</td><td>0.073</td><td>0.143</td><td>0.000</td><td>0.029</td><td>0.002</td><td>0.039</td><td>0.000</td><td>0.002</td></tr><tr><td>\(BraHab^{24} \)</td><td>0.048</td><td>0.015</td><td>0.179</td><td>0.052</td><td>0.054</td><td>0.139</td><td>0.008</td><td>0.007</td><td>0.001</td><td>0.017</td><td>0.076</td></tr><tr><td>\(ShaPl^{25} \)</td><td>0.085</td><td>0.087</td><td>0.044</td><td>0.161</td><td>0.016</td><td>0.011</td><td>0.004</td><td>0.018</td><td>0.000</td><td>0.198</td><td>0.044</td></tr><tr><td>\(ExRoPed^{26} \)</td><td>0.004</td><td>0.002</td><td>0.121</td><td>0.000</td><td>0.093</td><td>0.044</td><td>0.233</td><td>0.046</td><td>0.000</td><td>0.003</td><td>0.028</td></tr><tr><td>\(RoConst^{27} \)</td><td>0.003</td><td>0.035</td><td>0.082</td><td>0.179</td><td>0.016</td><td>0.009</td><td>0.048</td><td>0.054</td><td>0.000</td><td>0.025</td><td>0.103</td></tr><tr><td>\(RoSha^{28} \)</td><td>0.003</td><td>0.008</td><td>0.244</td><td>0.011</td><td>0.000</td><td>0.025</td><td>0.013</td><td>0.008</td><td>0.054</td><td>0.026</td><td>0.000</td></tr><tr><td>\(ExtColRo^{29} \)</td><td>0.001</td><td>0.022</td><td>0.428</td><td>0.000</td><td>0.135</td><td>0.003</td><td>0.081</td><td>0.032</td><td>0.009</td><td>0.049</td><td>0.001</td></tr><tr><td>ColRoPul30</td><td>0.120</td><td>0.001</td><td>0.051</td><td>0.036</td><td>0.006</td><td>0.038</td><td>0.008</td><td>0.017</td><td>0.038</td><td>0.051</td><td>0.239</td></tr><tr><td>ColRoCor31</td><td>0.117</td><td>0.020</td><td>0.041</td><td>0.005</td><td>0.148</td><td>0.004</td><td>0.111</td><td>0.009</td><td>0.036</td><td>0.151</td><td>0.092</td></tr><tr><td>TexRoEpi32</td><td>0.014</td><td>0.009</td><td>0.308</td><td>0.001</td><td>0.168</td><td>0.000</td><td>0.009</td><td>0.117</td><td>0.005</td><td>0.003</td><td>0.021</td></tr></table>
### c) Structure of the Germplasm Diversity
Factor 1 is positively correlated to the flowering and seed set ability and negatively correlated to the leaf colour, while factor 2 is positively correlated to the end branches colour, stem epidermis colour, leaf lobe margin, and the growth habit of stem (Figure 2). With regard to figure 2, four morphotypes were distinguished.
Morphotypes 1 and 2 exhibited good flowering and seed set ability but differed in terms of the end branches colour, stem epidermis colour, and growth habit of stem. The varieties belonging to morphotype 3 and 4 did not flower and differed in terms of the colour of the end branches, stem epidermis and the leaf lobe margin.
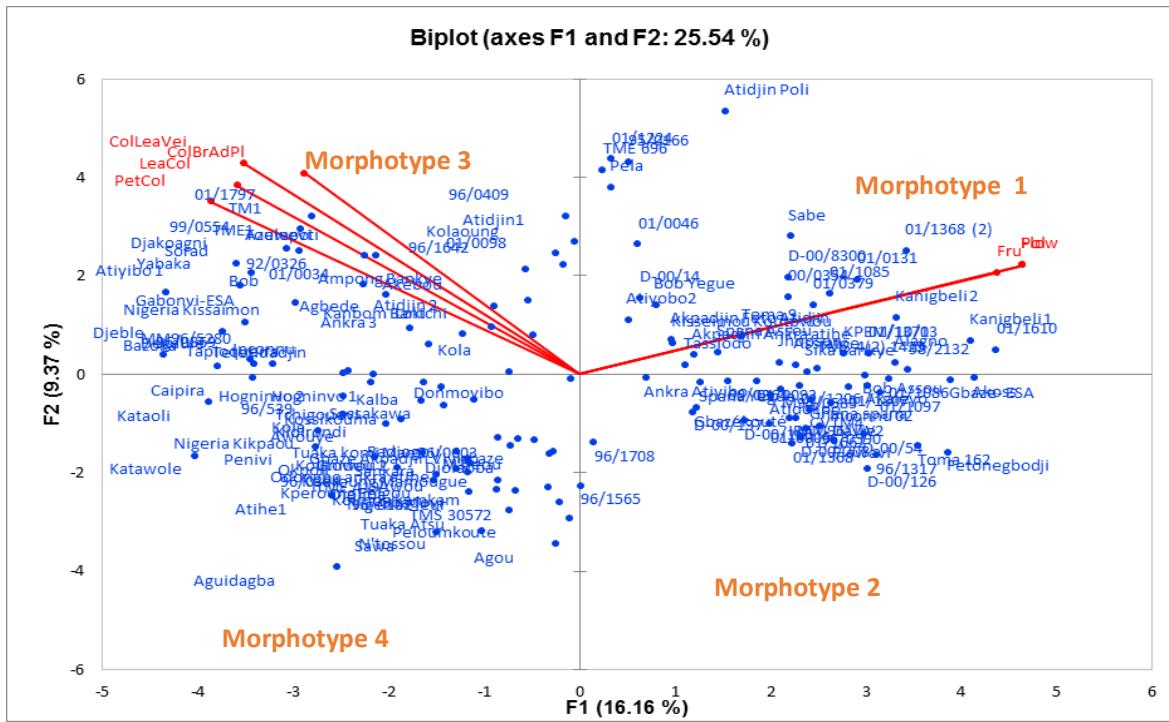 Figure 2: Component patterns of the MCA based upon factor 1 and 2 using significant phenotypic traits observed on 149 cassava cultivars of Togo
From the cluster analysis, the variance distribution function revealed that the optimal number of clusters was seven, with within class variance of 53.7 (Figure 3). Thus, the varieties were clustered into seven morphotypes (Figure 4). Morphotype 1 was composed of 137 varieties among which 53 were improved varieties, while remaining were landraces. Morphotype 1 is made of unflowering and unbranching cultivars with greenish-red petioles (Figure 4 and 5). Morphotype 2 was composed of 3 varieties (TMS 01_0379, Akoss, Kolaoung) exhibiting ovoid central leaflet, purple leaves, reddish green leaf vein, light green stem cortex, greyny-yellowish stem exterior, three levels of branching, dichotomous branching habit, conical root, cream root pulp, good flowering and seed set ability. Morphotype 3
The fourth morphotype comprised of cultivars Akebou, Akpadjin and Tassiodo is characterized by red petioles, obovate-lanceolate central leaflet, compact plants, pink root cortex, white root pulp and bad seed set hability; whereas cultivars belonging to morphotype 5 (Alagno, Pela) exhibited good leaf retention, irregularly shaped roots, many root constrictions, good flowering and seed set abilities (Figure 4 and 5). In morphotype 6, there were 8 cultivars (D00_126, Inconnu 2, D00_137, D00_208, D00_14, D00_54, D00_166 and D00_8300) with sessile peduncle root, conical cylindrical root, orange root pulp, short distance between leaf scars, good flowering and seed set ability. The morphotype 7 composed of cultivar TMS 96_0590 exhibited dark green apical leaves, cream stem epidermis, two levels of
branching, good flowering and seed set ability (Figure 4 and 5).
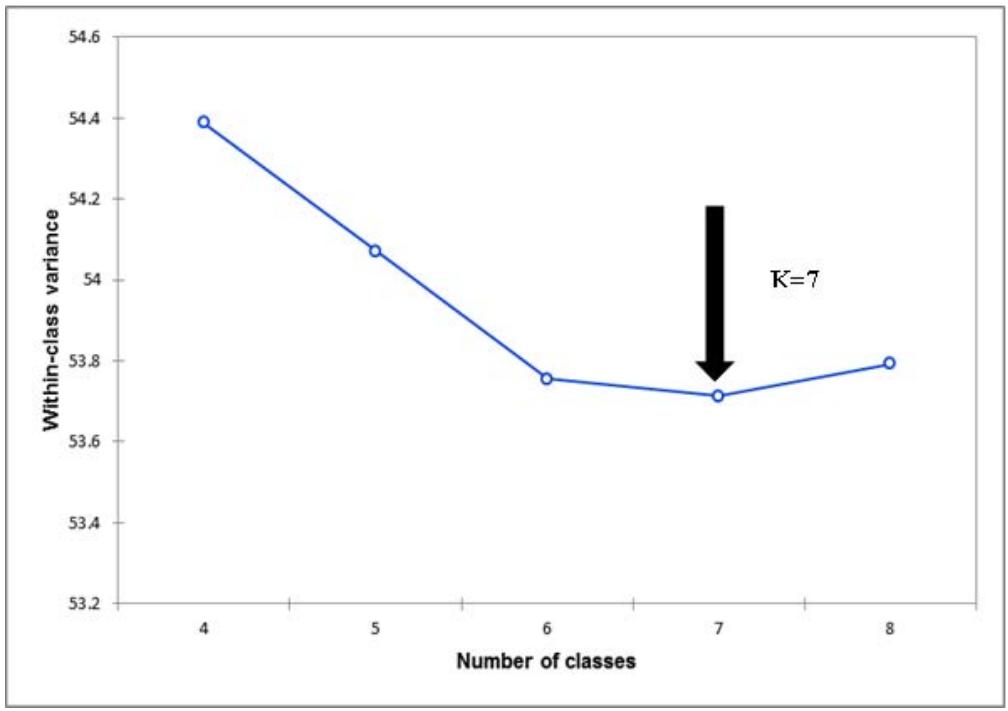 Figure 3: Distribution of the variance function according to the number of clusters obtained from cluster analysis based on significant morphological traits

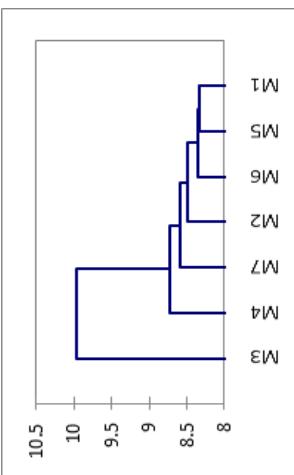
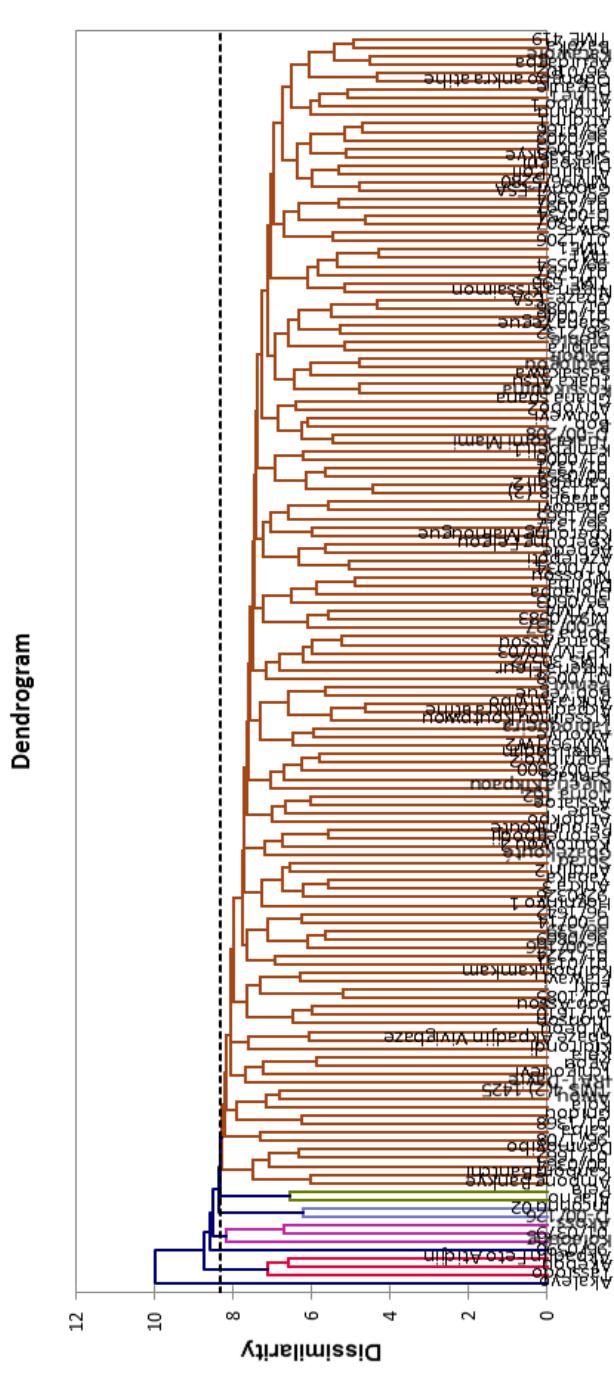 Figure 4: Dendrogram of 149 cassava genotypes revealed by the Wards method based on significant morphological
traits From left to right of the dendrogram the Morphotype 3, Morphotype 4, Morphotype 7, Morphotype 2, Morphotype 6, and Morphotype 5 are respectively represented.

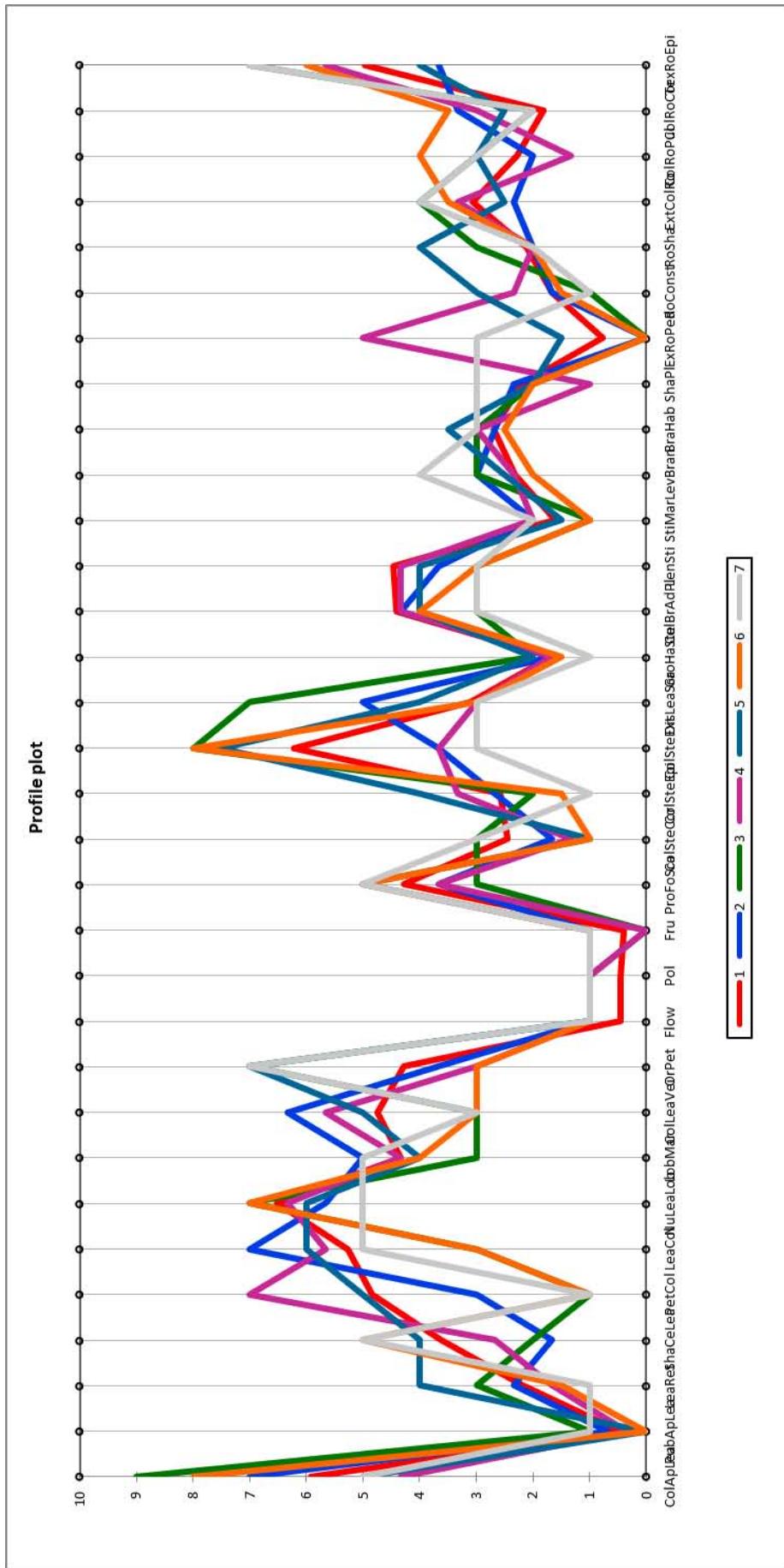 Figure 5: Profile plot describing the seven morphotypes obtained from the cluster analysis based on 32 morphological traits evaluated in 149 cassava cultivars of Togo
## IV. DISCUSSION
### a) Morphological Diversity of the Germplasm
Phenotyping of plant materials based on morphological traits has been used to determine the phenotypic variability among cultivars (Avijala et al., 2015; Agre et al., 2015; Adjebeng-Danquah & Gracen, 2020). The use of these traits allows rapid identification of cultivars. In addition, morphological traits are found to be stable, highly heritable and independent from the environment (Fukuda et al., 2010). However, molecular characterization may allow a more accurate detection of differences between germplasm bank cultivars than morphological characterization.
In our study, substantial variation was observed within the germplasm. Traits such as petiole colour, leaf colour, colour of leaf vein, flowering ability, seed set ability, levels of branching, colour of end branches, colour of stem epidermis, leaf lobe margins, growth habit of stem, and root flesh colour were underscored as the most relevant traits for cultivars discrimination. Genetic variability for morphological traits has been reported in different studies in Ghana (Asare et al., 2011; Adjebeng-Danquah & Gracen, 2017); in Benin (Agre et al.,2017), in Burkina Faso (Gmakouba et al., 2018) and in Brazil (Oliveira et al., 2015).
Root flesh colour is a trait with great importance for cassava because of dietary habits of Togolese. Moreover, this trait is directly related to the presence of vitamin. Orange varieties are rich carotenoids (provitamin A) (Kamanda et al., 2020). Low occurrence was found for yellow root colour and pink root colour varieties, which possibly have lycopene in their roots.
### b) Structure of the Germplasm Diversity
Cluster analysis classified the varieties into seven morphotypes, showing random distribution of the varieties. The fact that cassava is an outcrossing crop which can propagate vegetatively could explain this result. This facilitates the dispersion of varieties the exchange among farmers and, consequently, the exchange of genes (Agre et al., 2017). The main factor involved in the high diversity found may be gene flow promoted by farmers, who have acted as a dispersing agents for the species. An intense exchange system of varieties has been documented among farmers growing cassava.
The germplasm bank cultivars were not grouped based on the geographical origin distribution. Cultivars collected from place such as Vogan, Wetrope, Akebou, Danyi, Aouda, Davie, and Assoukoko were clustered in morphotype 1. Likewise cultivars from Bafilo, Assoukoko, Danyi, and Bourondè were also clustered together in morphotype 2. The remaining clusters also included cultivars from different collection regions. The informal farmers to farmers seed supply system practiced in the country could explained this result. This agrees with earlier studies on cassava (Ojulong et al., 2010; Sing et al., 2015; Adjebeng-Danquah & Gracen, 2017, Gmakouba et al., 2018). In addition, the cassava cultivars collected from the same region were clustered into different morphotypes which suggest a high genetic diversity within each collection area. Similar findings were also reported by Agre et al. (2017) in Bénin. Moreover, there was no clear differentiation and real structuring between local and improved varieties in this study as also reported by Kombo et al. (2012).
Morphotypes identified may be valuable in cassava germplasm management and cultivars identification. Especially, the varieties belonging to morphotypes 5, 6 and morphotype 7 might be most desirable for breeding due to their good flowering and seed set ability and adaptability to environmental conditions. These cultivars could be used to set up crossing blocks in order to develop segregating breeding lines with farmers desired traits.
## V. CONCLUSIONS
The study revealed that the cassava germplasm of Togo exhibited high phenotypic diversity. Morphological traits such as petiole colour, leaf colour, colour of leaf vein, flowering ability, seed set ability, levels of branching, colour of end branches, colour of stem epidermis, leaf lobe margins and the growth habit of stem were the most diverse and could be used for cultivar identification in the field. The varieties of morphotypes 5, 6 and 7 harboured interesting features such as flowering and seed set ability and may be usefull for the national breeding programme. For breeding purpose, superior parental clones could be selected from these morphotypes for crossing and generating a breeding population.
### ACKNOWLEDGEMENTS
The authors are thankful to farmers and the agricultural extension agents who participated in the germplasm collection, as well as the Togolese Agronomic Research Institute (ITRA-CRAL) for offering the logistical assistance.
Funding: This manuscript is extracted from PhD. thesis, which was co-funded by ECONET-Foundation and the Germany Academic Exchange Service (WACCI/DAAD) at West Africa Centre for Crop Improment (University of Ghana).
Authors' Contributions: GMAKOUBA Tighankoumi carried out the study, analyzed the data and drafted the manuscript. DZIDZIENYO K. Daniel, SOME Koussao, TONGOONA Pangirayi and ASANTE I. Kwame participated in the study design and were major contributors in writing and correcting the manuscript. All authors read and approved the final manuscript.
Consent for Publication: The authors declare that they obtained an informed consent for publication from people involved in this study.
Availability of Data and Materials: Data are within the paper and its supporting information files. The datasets are fully available without restriction on reasonable request from the corresponding author.
Competing Interests: The authors declare that they have no competing interests.
Generating HTML Viewer...
References
26 Cites in Article
Joseph Adjebeng-Danquah,Vernon Gracen (2017). Agronomic Performance and Genotypic Diversity for Morphological Traits among Cassava Genotypes in the Guinea Savannah Ecology August.
Joseph Adjebeng-Danquah,Joseph Manu-Aduening,Isaac Asante,Richard Agyare,Vernon Gracen,Samuel Offei (2020). Genetic diversity and population structure analysis of Ghanaian and exotic cassava accessions using simple sequence repeat (SSR) markers.
A Agre,R Bhattacharjee,I Y Rabbi,O A Alaba,N N Unachukwu,M Ayenan,Y L Loko,G J Bauchet,A Dansi (2017). Classification of elite cassava varieties (Manihot esculenta Crantz) cultivated in Benin Republic using farmers’ knowledge, morphological traits and simple sequence repeat (SSR) markers.
Matoso Avijala,Leonardo Francisco,Leonardo Bhering,De Azevedo,Damião Peixoto,Pedro Cruz,Souza Crescêncio,Constantino Carneiro,Anabela Estevão Cuambe,Zacarias (2015). Evaluation of Cassava Genotypes Reveals Great Genetic Variability and Potential Selection Gain.
A Banito,K Kpémoua,K Wydra (2010). Screening of Cassava Genotypes for Resistance to Bacterial Blight Using Strain x Genotype Interactions.
Hernan Ceballos,Juan Perez,Orlando Joaqui-Barandica,Jake Lenis,Morante,F Nelson & Calle,Lizbeth Pino,Clair Hershey (2016). Cassava Breeding I: The Value of Breeding Value.
T Costa,O Vidigal Filho,M Vidigal,M Galván,G Lacanallo,Silva Li,And Kvitschal,M (2013). Genetic diversity and population structure of sweet cassava using simple sequence repeat (SSR) molecular markers.
K Ezui (2017). Understanding the productivity of cassava in West Africa.
Ifad Fao,Unicef Who (2023). The State of Food Security and Nutrition in the World 2023.
João Giles,D Antonio,Gleison Oliosi,P Weverton,Heder Rodrigues,Ana Braun,Fábio Ribeiro-Barros,Partelli (2018). Agronomic Performance and Genetic Divergence between Genotypes of Manihot Esculenta.
Tighankoumi Gmakouba,Somé Koussao,Ernest Traore,Kossi Kpemoua,Jean-Didier Zongo (2018). Analyse de la diversité agromorphologique d’une collection de manioc (<i>Manihot esculenta</i> Crantz) du Burkina Faso.
C Guevara,R Kawuki,M E Ferguson,I Kamanda,E Blay,I Asante (2020). Selected Morphological and Agronomic Descriptors for the Characterization of Cassava.
R Kawuki,M Ferguson,M Labuschagne,L Herselman,J Orone,I Ralimanana,M Bidiaka,S Lukombo,M Kanyange,G Gashaka,G Mkamilo,J Gethi,H Obiero (2011). Variation in qualitative and quantitative traits of cassava germplasm from selected national breeding programmes in sub-Saharan Africa.
G Kombo,A Dansi,L Loko,G Orkwor,R Vodouhè,P Assogba,J Magema (2012). Diversity of cassava (Manihot esculenta Crantz) cultivars and its management in the department of Bouenza in the Republic of Congo.
Henry Ojulong,Maryke Labuschagne,Liezel Herselman,Martin Fregene (2010). Yield traits as selection indices in seedling populations of cassava.
Bright Peprah,Elizabeth Parkes,Joseph Manu-Aduening,Peter Kulakow,Angeline Van Biljon,Maryke Labuschagne (2020). Genetic variability, stability and heritability for quality and yield characteristics in provitamin A cassava varieties.
Wirulda Pootakham,Jeremy Shearman,Panthita Ruang-Areerate,Chutima Sonthirod,Duangjai Sangsrakru,Nukoon Jomchai,Thippawan Yoocha,Kanokporn Triwitayakorn,Somvong Tragoonrung,Sithichoke Tangphatsornruang (2014). Large-Scale SNP Discovery through RNA Sequencing and SNP Genotyping by Targeted Enrichment Sequencing in Cassava (Manihot esculenta Crantz).
Ismail Rabbi,Peter Kulakow,Joseph Manu-Aduening,Ansong Dankyi,James Asibuo,Elizabeth Parkes,Tahirou Abdoulaye,Gezahegn Girma,Melaku Gedil,Punna Ramu,Byron Reyes,Mywish Maredia (2015). Tracking crop varieties using genotyping-by-sequencing markers: a case study using cassava (Manihot esculenta Crantz).
Ismail Rabbi,Martha Hamblin,P Kumar,Melaku Gedil,Andrew Ikpan,Jean-Luc Jannink,Peter Kulakow (2014). High-resolution mapping of resistance to cassava mosaic geminiviruses in cassava using genotyping-by-sequencing and its implications for breeding.
D Rogers,S Appan (1973). Manihot and Manihotoides (Euphorbiaceae). A computerassisted study.
Selamawit Abebe,Danga Gitore,Henga Benjamin,Gurmu Sylvia,Fekadu (2021). Phenotypic Characterization of Sweet Potato Genotypes in Ethiopia for Selection of Those Possessing Optimal Dual-Purpose.
Geoffrey Sing,D Ateka,S Miano,Theresia Githiri,Munga (2015). Assessment of the Responses of Cassava Breeder ' s Germplasm to Cassava Mosaic Virus Infection in Kenya.
Marcos Siqueira,Aline Borges,Teresa Valle,Elizabeth Veasey (2011). A comparative genetic diversity assessment of industrial and household Brazilian cassava varieties using SSR markers.
Jean Sogbedji,Lakpo Mianikpo,Kodjovi Kokou Agboyi,Ruth Sotomè Detchinli,Mihikouwe Atchoglo,Mazinagou (2015). Sustaining Improved Cassava Production on West African Ferralsols Through Appropriate Varieties and Optimal Potassium Fertilization Schemes.
Monique Soro,Serge Marie Felicien Wend-Pagnagdé Zida,Koussao Somé,Fidèle Tiendrébéogo,Daniel Otron,Justin Pita,James Néya,Daouda Koné (2024). Estimation of Genetic Diversity and Number of Unique Genotypes of Cassava Germplasm from Burkina Faso Using Microsatellite Markers.
Á Vidal,L Vieira,C Ferreira,F Souza,A Souza,C Ledo (2015). Genetic fidelity and variability of micropropagated cassava plants (Manihot esculenta Crantz) evaluated using ISSR markers.
Explore published articles in an immersive Augmented Reality environment. Our platform converts research papers into interactive 3D books, allowing readers to view and interact with content using AR and VR compatible devices.
Your published article is automatically converted into a realistic 3D book. Flip through pages and read research papers in a more engaging and interactive format.
Our website is actively being updated, and changes may occur frequently. Please clear your browser cache if needed. For feedback or error reporting, please email [email protected]
Thank you for connecting with us. We will respond to you shortly.