Antimicrobial peptides (AMPs) are bioactive molecules known for their strong antimicrobial properties against a variety of microorganisms. The increasing prevalence of antibiotic-resistant infections, coupled with a scarcity of newly developed antibiotics, has intensified the search for novel sources of AMPs. Plant-derived AMPs present several benefits compared to conventional antibiotics. Consequently, this research focused on screening various plant species to identify new antimicrobial peptides. Water-soluble proteins were extracted from the dried plant materials using an aqueous extraction buffer. The resulting extracts were evaluated for antibacterial activity, with the minimum inhibitory concentration (MIC) being determined, followed by separation through thin layer chromatography and subsequent agar overlay bioautography. Protein extracts from five different plants exhibited antibacterial properties against both Gram-positive and Gram-negative bacteria, with inhibition zones measuring between 11 and 22 mm.
## I. INTRODUCTION
Every year, around 700,000 patients succumb to antimicrobial resistance (AMR) globally. Projections indicate that this mortality rate could rise to 10 million by the year 2050 (O'Neill, 2014).
The emergence of superbugs, which are pathogenic bacteria resistant to the majority or entirety of available antibiotics, presents a significant challenge. The World Health Organization has cautioned that these multidrug-resistant pathogens could potentially revert global health to conditions reminiscent of the preantibiotic era (WHO, 2017).
There is a pressing necessity to identify novel alternatives to address the issue of antimicrobial resistance. Potential solutions include bacteriophages, antibodies, probiotics, lysins, and antimicrobial peptides (AMPs), which have shown promise in tackling this critical challenge (Ghosh, Sarkar, Issa, & Haldar, 2019).
AMPs are short oligopeptides, comprising up to 50 amino acids, that are present in all living organisms. Plant-derived AMPs represent a valuable natural alternative to synthetic antibiotics for applications in human healthcare and agriculture, particularly for protection and healing (Almeida et al., 2020; Tang, Prodhan, Biswas, Le, & Sekaran, 2018).
These peptides exhibit a wide range of activities, including antibacterial, antifungal, antiviral, and even anticancer properties (R. E. W. Hancock, 2001; Kamysz, Okrój, & Lukasiak, 2003). Their mechanisms of action are varied and closely linked to their structural characteristics; AMPs interact electrostatically with bacterial cell walls, leading to a disruption of membrane integrity (Tzong-Hsien, Kristopher, & Marie-Isabel, 2016). Additionally, these peptides possess the ability to penetrate biological membranes, allowing them to exert effects intracellularly (Cardoso et al., 2019). Beyond their antimicrobial functions, AMPs also play a significant role in immunomodulation (Haney & Hancock, 2013).
The broad activity spectrum, straightforward synthesis, and specific mechanisms of action associated with antimicrobial peptides (AMPs) significantly mitigate the risk of bacterial resistance. This makes them attractive candidates for the development of new therapeutic agents to address the pressing issue of bacterial resistance (Baltzer & Brown, 2011). There is a critical need to identify new AMPs that can either replace or improve the effectiveness of current antibiotics. Moreover, AMPs can be easily obtained from natural sources, such as plants, for applications in human healthcare, agriculture, and the food industry.
This research focused on screening a variety of plant species to identify novel antimicrobial peptides. We achieved this by extracting protein and peptide fractions from the plants and evaluating their antimicrobial efficacy against several standard bacterial strains. The study involved determining the minimum inhibitory concentration of the extracts exhibiting antibacterial properties, isolating the plant protein mixture into distinct proteins and peptide fractions, and localizing the active compounds present in the extracts with antibacterial activity.
## II. MATERIALS AND METHODS
### a) Plant Material
Seeds and leaves of various plants screened for the presence of antimicrobial peptides (AMPs) are listed in Table 1.
Table 1: List of Various Plants Screened for the Presence of Antimicrobial Peptides
<table><tr><td>No.</td><td>Binomial Name</td><td>Family</td><td>Part tested</td><td>Protein Concentration (μg/ml)</td></tr><tr><td>1.</td><td>Punica granatum</td><td>Lythraceae*</td><td>seeds</td><td>261</td></tr><tr><td>2.</td><td>Vitis labrusca 'Fragola'</td><td>Vitaceae*</td><td>seeds</td><td>4064</td></tr><tr><td>3.</td><td>Annona</td><td>Annonaceae*</td><td>seeds</td><td>4285</td></tr><tr><td>4.</td><td>Ricinus communis</td><td>Euphorbiaceae*</td><td>seeds</td><td>1797</td></tr><tr><td>5.</td><td>Ocimum basilicum</td><td>Lamiaceae*</td><td>seeds</td><td>4561</td></tr><tr><td>6.</td><td>Foeniculum vulgare</td><td>Apiaceae*</td><td>Grounded seeds</td><td>2557</td></tr><tr><td>7.</td><td>Cucumis melo flexuosus</td><td>Cucurbitaceae*</td><td>seeds</td><td>5806</td></tr><tr><td>8.</td><td>Ammi visnaga</td><td>Apiaceae*</td><td>seeds</td><td>1037</td></tr><tr><td>9.</td><td>Brassica rapa subsp. Rapa</td><td>Brassicaceae*</td><td>seeds</td><td>1659</td></tr><tr><td>10.</td><td>Matricaria recutita</td><td>Asteraceae*</td><td>Leaves, flowers</td><td>2351</td></tr><tr><td>11.</td><td>Raphanus sativus</td><td>Brassicaceae*</td><td>seeds</td><td>2156</td></tr><tr><td>12.</td><td>Coriandrum sativum</td><td>Apiaceae*</td><td>Leaves</td><td>2557</td></tr><tr><td>13.</td><td>Lepidium sativum</td><td>Brassicaceae*</td><td>Leaves</td><td>414</td></tr><tr><td>14.</td><td>Thymus vulgaris</td><td>Lamiaceae*</td><td>Leaves</td><td>3253</td></tr><tr><td>15.</td><td>Olea europaea</td><td>Oleaceae*</td><td>Leaves</td><td>4936</td></tr><tr><td>16.</td><td>Psidium guajava</td><td>Myrtaceae*</td><td>Leaves</td><td>2323</td></tr><tr><td>17.</td><td>Zingiber officinale</td><td>Zingiberaceae*</td><td>roots</td><td>2653</td></tr><tr><td>18.</td><td>Pimpinella anisum</td><td>Apiaceae*</td><td>Leaves</td><td>3285</td></tr><tr><td>19.</td><td>Rosmarinus officinalis</td><td>Lamiaceae*</td><td>Leaves</td><td>1543</td></tr><tr><td>20.</td><td>Trigonella foenum-graecum</td><td>Fabaceae*</td><td>Leaves</td><td>4147</td></tr><tr><td>21.</td><td>Nigella sativa</td><td>Ranunculaceae *</td><td>seeds</td><td>4117</td></tr><tr><td>22.</td><td>Rumex obtusifolius L.</td><td>Polygonaceae*</td><td>seeds</td><td>1707</td></tr><tr><td>23.</td><td>Salvia hispanica</td><td>Lamiaceae*</td><td>seeds</td><td>1102</td></tr><tr><td>24.</td><td>Cuminum cyminum</td><td>Apiaceae*</td><td>Grounded seeds</td><td>1382</td></tr><tr><td>25.</td><td>Cinnamomum zeylanicum</td><td>Lauraceae*</td><td>Cinnamon sticks</td><td>2599</td></tr><tr><td>26.</td><td>Prunus dulcis</td><td>Rosaceae*</td><td>seeds</td><td>1974</td></tr><tr><td>27.</td><td>Hibiscus sabdariffa</td><td>Malvaceae*</td><td>flowers</td><td>1828</td></tr><tr><td>28.</td><td>Brassica oleracea var. capitata f. rubra</td><td>Brassicaceae*</td><td>Leaves</td><td>4212</td></tr></table>
### b) Bacteria, Growth Media and Reagents
The isolated antimicrobial peptides (AMPs) were evaluated for their efficacy against various bacterial strains from the ATCC collection, which included Gram- positive species such as Staphylococcus aureus ATCC 12493, Streptococcus epidermidis ATCC 12228, and Bacillus subtilis ATCC 6633, as well as Gram-negative species including Escherichia coli NCTC 13846,
Klebsiella pneumoniae ATCC 2021, Pseudomonas aeruginosa ATCC 27853, and Acinetobacter baumannii ATCC 19606.
Muller Hinton Agar (MHA), Nutrient Agar (NA), Nutrient Broth (NB), Brain Heart Infusion Broth (BHIB), 50 mM phosphate buffer pH 7, 2 mM EDTA, Glycerol, 50 mM NaCl, distilled water (D.W), tap water, 0.1% 2,3,5 triphenyl tetrazolium chloride (TTC), n-butanol, acetone, acetic acid, 5% ammonia, ninhydrin, methylthiazol tetrazolium (MTT).
### c) Total Water-Soluble Proteins Extraction
The plant material underwent a washing process with tap water followed by distilled water, after which it was dried under sunlight. For the extraction of water-soluble proteins, the milled plant material was combined with a cold extraction buffer at a ratio of 1:10 (w/v), consisting of $50~\mathrm{mM}$ phosphate buffer at pH 7, 2 mM EDTA, $5\%$ glycerol, and $50~\mathrm{mM}$ NaCl. Subsequently, the mixture was agitated on a shaker for two hours at a temperature of $4^{\circ}\mathrm{C}$, followed by centrifugation at $12,000~\mathrm{rpm}$ for 20 minutes at the same temperature. The clear solution was then filtered through sterile gauze and stored at $-20^{\circ}\mathrm{C}$ (Aliahmadi, Roghanian, Emtiazi, & Ghassemi pour, 2011).
### d) Total Protein Concentration Estimation
The concentration of the extracted water-soluble proteins was estimated using a nano-drop spectrophotometer.
### e) Testing Extracts on Bacteria by Agar Well Diffusion Assay
Muller Hinton agar (MHA) plates were inoculated using a cotton swab that had been moistened with a McFarland standardized test organism. Using a sterile pipette blue tip, holes measuring 6 to $8\mathrm{mm}$ in diameter were created and subsequently filled with $50~\mu \mathrm{L}$ of the plant extract. The petri dishes were then incubated at $37^{\circ}\mathrm{C}$ for a duration of 24 hours. The diameters of the zones of growth inhibition were measured. The inoculum was prepared in a sterile saline solution, with its turbidity adjusted to the 0.5 McFarland standard $(10^{8}\mathrm{CFU / mL})$. A $3\%$ potassium iodide (KI) solution was utilized as a positive control (Abed, 2015; Nigussie, Davey, Legesse, Fekadu, & Makonnen, 2021).
### f) Determination of Minimum Inhibitory Concentration (MIC) by Microtiter Broth Dilution Method
In a 96-well microtiter plate, $100~\mu \mathrm{L}$ of plant extract was introduced into the first well, while $50~\mu \mathrm{L}$ of BHIB media was allocated to each of the remaining wells. Subsequently, $50~\mu \mathrm{L}$ was extracted from the first well and transferred to the second well, continuing this process until the last well to achieve serial dilution for each extract. A volume of $50~\mu \mathrm{L}$ of bacterial suspension, calibrated to the 0.5 McFarland standard, was added to each well, with the exception of the last well designated as the "negative control." The plates were then covered and incubated at $37^{\circ}\mathrm{C}$ for 24 hours. Following this incubation, $20~\mu \mathrm{L}$ of $0.1\%$ 2,3,5 triphenyl tetrazolium chloride (TTC) was introduced into each well, and the plates were re-incubated for an additional 15 minutes. The plates were subsequently examined, and the final dilution that exhibited antibacterial activity was identified by noting the last well that did not display red coloration, as the presence of red color indicates bacterial growth (Abou-Elkhair, Fadda, & Abu-Mohsen, 2010).
### g) Protein and Peptide Separation by Thin Layer Chromatography (TLC)
This technique was derived from the work of Jaskiewicz et al. (2016), utilizing Silica gel 60 RP-18 F254S (Merck) thin-layer chromatography (TLC) plates as the stationary phase. A volume of $5\mu \mathrm{L}$ from each plant extract was applied to the TLC plate. The separation process employed a mobile phase composed of n-butanol, acetone, acetic acid, $5\%$ ammonia, and distilled water in a ratio of 4.5:1.5:1:1:2 (v/v/v/v/v) within a sealed chamber. The separation was allowed to proceed until the solvent front reached the end line, at which point the plates were removed and air-dried to facilitate the evaporation of the eluents. Subsequently, the dried plates were treated with a $2\%$ (w/v) ninhydrin solution in a mixture of acetone and glacial acetic acid (25:1, v/v). Following this, the plates were left to dry for several minutes at ambient temperature before being heated in an oven at $80^{\circ}\mathrm{C}$ until the peptide zones became visible (Gwarda, Tomczyszyn, Misicka, & Dzido, 2013; Sharma, Abid, & Sajgotra, 2017). For subsequent antibacterial assays, the TLC plates were sterilized under ultraviolet light for 15 minutes. The identification of specific compounds was constrained by the lack of available reference standards.
### h) TLC-Agar-Overlay Bioautography
Developed TLC plates were placed within a sterile Petri dish. A suspension of representative bacterial strains, measuring one milliliter, was combined with $10~\mathrm{mL}$ of melted Mueller-Hinton agar, which was then poured as a thin layer over the developed TLC plate. Following the solidification of the medium, the TLC plate was incubated for 24 hours at a temperature of $35 \pm 2^{\circ}\mathrm{C}$. Subsequently, the TLC-bioautography plates were sprayed with an aqueous solution of methylthiazol tetrazolium (MTT) at a concentration of $2.5~\mathrm{mg/mL}$ (Sigma, USA). Clear zones of inhibition were noted against a purple background (Valle, Puzon, Cabrera, & Rivera, 2016).
## III. RESULTS
### a) Total Protein Concentration Estimation
The plant protein extracts examined in this study exhibited a range of protein concentrations between 261 and $5806\mu \mathrm{g / ml}$. Among these, the protein extract from
Punica granatum displayed the lowest protein content, while the extract from Cucumis melo flexuosus demonstrated the highest protein concentration. Detailed protein concentrations for each plant extract can be found in Table (1).
### b) Testing Plants Protein Extracts on Bacteria by Agar Well Diffusion Assay
The protein extracts derived from five different plant species exhibited varying levels of antibacterial activity against a range of both Gram-positive and Gram-negative bacteria, with inhibition zones measuring between 11 and $22\mathrm{mm}$. Extracts from Ricinus communis and Vitis labrusca demonstrated antibacterial effects against the Gram-positive bacteria S. aureus and S. epidermidis. The extract from Raphanus sativus displayed a more extensive antibacterial spectrum, affecting both Gram-positive (S. aureus and S. epidermidis) and Gram-negative bacteria (E. coli and A. baumannii). Notably, none of the protein extracts exhibited antibacterial activity against B. subtilis or K. pneumoniae. Antibacterial effects were recorded against S.aureus for the extracts of R. sativus, R. communis, V. labrusca, and Prunus dulcis, with inhibition zones measuring 16, 15, 18, and $14\mathrm{mm}$, respectively (Figure 1).
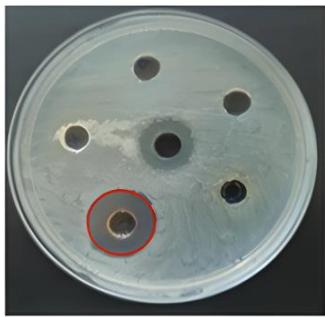 (A)
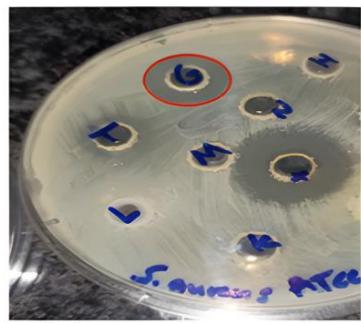 (B)
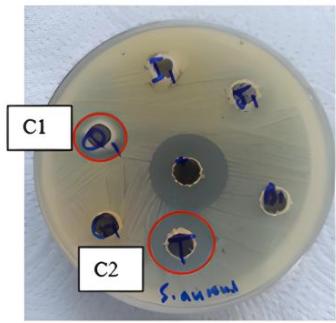 (C) Figure 1: Antibacterial activity of plant protein extracts against S. aureus(A) Vitis labrusca, (B) Ricinus communis, (C1) Prunus dulcis and (C2) Raphanus sativus
Figure 2 illustrates that the extracts of $R.$ sativus, $R.$ communis, and $V.$ labrusca exhibit antibacterial properties against S. epidermidis, resulting in zones of inhibition measuring 20 mm, 22 mm, and 11 mm, respectively.
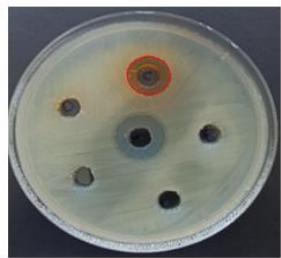 (A)
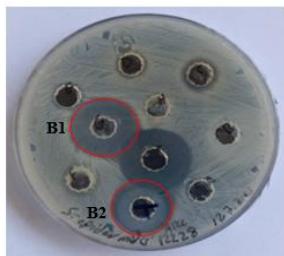 (B) Figure 2: Antibacterial activity of plant protein extracts against S. epidermidis.(A) Vitis labrusca, (B1) Ricinus communis, (B2) Prunus dulcis
R. sativus has demonstrated antibacterial properties against E. coli, exhibiting an inhibition zone of $19\mathrm{mm}$ and against A. baumannii, with an inhibition zone of $16\mathrm{mm}$, as illustrated in Figure 3.
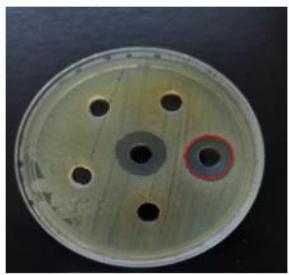 (A)
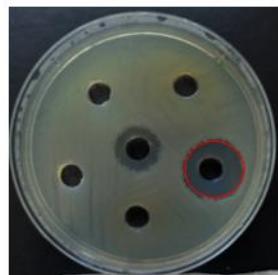 (B) Figure 3: Antibacterial activity of R. sativus protein extract. (A) A. baumannii and (B) E. coli
The growth of $P$ aeruginosa was suppressed by the protein extract of $H$ sabdariffa, resulting in a modest zone of inhibition measuring $12\mathrm{mm}$, as illustrated in Figure 4.
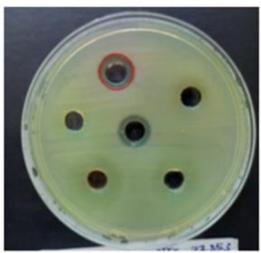 Figure 4: Antibacterial Activity of Hibiscus Sabdariffaprotein Extract Against P. Aeruginosa.
### c) Determination of Minimum Inhibitory Concentration (MIC) by Microtiter Broth Dilution Method
The minimum concentration that resulted in the inhibition of bacterial growth by plant protein extracts was determined through the observation of color changes in the wells of a 96 microtiter plate, utilizing the micro broth dilution assay alongside TTC as a growth indicator. The protein extracts from V. labrusca, R. communis, R. sativus, and P. dulcis exhibited growth inhibition against S. aureus at concentrations of 2.03, 0.45, 0.54, and $1.97\mathrm{mg / ml}$, respectively. Conversely, the protein extracts from V. labrusca, R. communis, and R. sativus demonstrated inhibitory effects on the growth of S. epidermidis at concentrations of 4.06, 0.45, and $0.54\mathrm{mg / ml}$. Additionally, the growth of A. baumannii and E. coli was inhibited at concentrations of $1.07\mathrm{mg / ml}$ and $0.53\mathrm{mg / ml}$ for R. sativus, respectively. Furthermore, the growth of P. aeruginosa was inhibited by a concentration of $0.45 \mathrm{mg} / \mathrm{ml}$ of $H.$ sabdariffa protein extract.
### d) Protein and peptide separation by Thin Layer Chromatography (TLC)
Antibacterial activity from plant protein extracts was isolated using the thin-layer chromatography (TLC) technique, and the resulting fractions were analyzed with ninhydrin spray. The separation of extracts from V. labrusca (Fig. 5-A), P. dulcis (Fig. 5-D), and P. guajava (Fig. 5-E) yielded two distinct bands. In contrast, the separation of R. communis resulted in a continuous line featuring two distinct bands, one positioned at the beginning and the other at the end of the migration line (Fig. 5-B). The separation of Raphanus sativus displayed two intertwined bands (Fig. 5-C). Additionally, the separation of H. sabdariffa resulted in three distinct bands.
 (A)
 (B)
 (C)
 (D)
 (E) Figure 5: TLC plates of plant protein extracts visualized after coloring with ninhydrin solution. (A) Vitis labrusca, (B) Ricinus communis, (C) Raphanus sativus, (D) Prunus dulcis, and (E) Psidium guajava. Arrows point to separated fractions
### e) TLC-Agar-Overlay Bioautography
The antibacterial activity of the compounds separated on TLC was further evaluated using thin-layer chromatography (TLC) agar overlay bioautography. The antibacterial properties of V. labrusca, R. communis, R. sativus, Prunus dulcis, Psidium guajava, and H. sabdariffa were evidenced by the presence of clear zones of inhibition against the respective bacteria on a purple background. Notably, V. labrusca, R. communis, and R. sativus exhibited inhibitory effects against both S. aureus and S.epidermidis. Additionally, R. sativus demonstrated inhibitory activity against E. coli and A. baumannii, while H. sabdariffa showed effectiveness against P. aeruginosa. Conversely, Prunus dulcis did not
produce any zones of inhibition against S. aureus. The antibacterial effects of V. labrusca (Fig. 6-A) and R. sativus (Fig. 6-C) against S.aureus are illustrated by clear zones near the origin line, with the absence of stained bands on the corresponding TLC plates indicating that the inhibitory compounds are non-protein in nature. Two stained compounds on the TLC plates of $R$ communis (Fig. 6-B) resulted in two zones of inhibition, one located near the origin line ( $R_{f} = 0.06$ ) and the other positioned in the middle of the TLC plate ( $R_{f} = 0.73$ ).
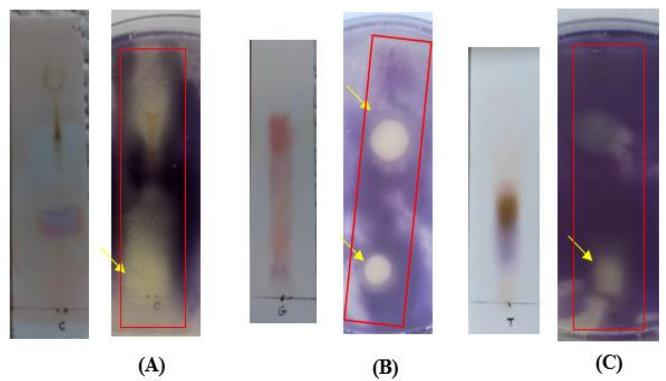 Figure 6: TLC-agar-overlay bioautography of seed extract of (A) V. labrusca, (B) R.communis, and (C) R. sativus against S. aureus. Growth inhibition zones are indicated by arrows.
Comparable outcomes were observed in agar overlay bioautography using S. epidermidis, where V. labrusca (Fig. 7-A) and R. sativus (Fig. 7-C) exhibited minor inhibitory zones near the origin line. In contrast, two stained compounds on the R. communis TLC plate (Fig. 7-B) revealed two distinct zones: one located close to the origin line ( $R_{f} = 0.06$ ) and the other positioned in the middle of the TLC plate ( $R_{f} = 0.73$ ).
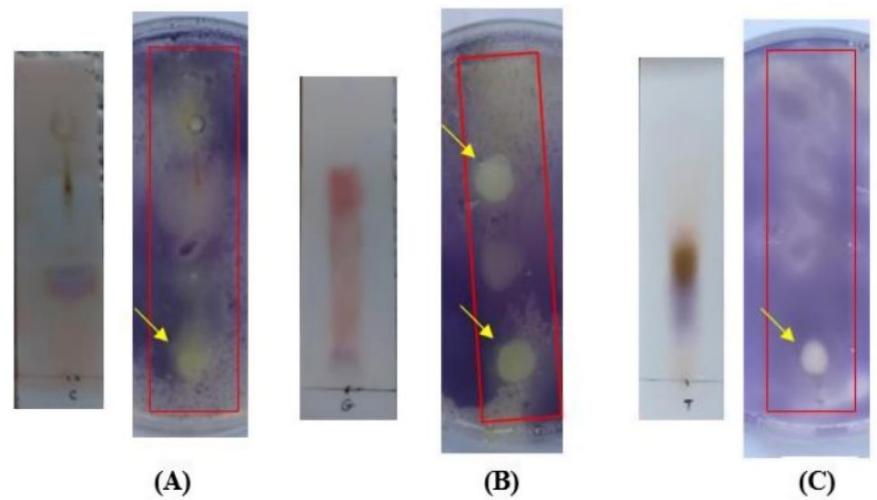 Figure 7: TLC-agar-overlay bioautography of seed extract of (A) V. labrusca, (B) R. communis, and (C) R. sativus against S. epidermidis
Non-stained fraction on the line of origin of $R$. sativus TLC plate exhibited a large inhibitory zone against E. coli (Fig. 8-A) and A. baumannii (Fig. 8-B).

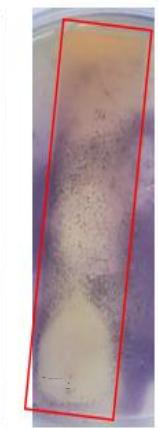 (A)

 (B) Figure 8: TLC-agar-overlay bioautography of R. sativus seeds extract against (A) E. coli NCTC 13846 and (B) A. baumannii ATCC 19606.
Figure 9 illustrates that the TLC plate of $H$ sabdariffa exhibited three inhibitory zones against $P$ aeruginosa. The first zone had an $R_{f}$ value of 0.11, the second zone had an $R_{f}$ value of 0.46, and the third, which was a non-stained compound, generated a zone close to the end line.

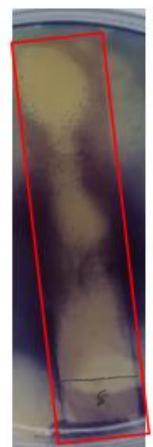 Figure 9: TLC-agar-overlay bioautography of $H.$ sabdariffa flowers extract against $P.$ aeruginosa
## IV. DISCUSSION
The rise in antimicrobial resistance to existing antibiotics is a major global health issue, limiting treatment options. This problem is worsened by the scarcity of newly developed antibiotics. Research shows that antibiotic resistance has increased worldwide over the past twenty years, mainly due to overprescription and misuse of antibiotics. Finding new antimicrobial agents is crucial, yet the development of new antibiotics has slowed down, stressing the need for innovative solutions (Klein et al., 2018; O'Neill, 2016).
Plants have been used for medicinal purposes for a long time and are important in discovering new antimicrobial agents. Many plants contain natural compounds that fight infections. Research emphasizes the value of plants as a source of new antimicrobial compounds due to their diverse secondary metabolites and cost-effectiveness (Cowan, 1999b; Newman & Cragg, 2012; Newman & Cragg, 2020; Atanasov et al., 2015). The extraction of antimicrobial compounds from plants has advantages over synthetic methods, such as lower toxicity risks and unique mechanisms of action. Therefore, exploring plants for new antimicrobial compounds can lead to effective treatments for drug-resistant infections (Anand et al., 2022; Akinyemi, Oladapo, Okwara, Ibe, & Fasure, 2005; Djeussi et al., 2013; Jubair, Rajagopal, Chinnappan, Abdullah, & Fatima, 2021).
Plants contain a diverse range of antimicrobial peptides that can be extracted and studied for their potential therapeutic applications (Broekaert et al., 1997;
Lei et al., 2019; Tam, Wang, Wong, & Tan, 2015). Several studies found that antimicrobial peptides were effective against drug-resistant bacteria, suggesting their potential as a therapeutic option for these infections (Chung & Khanum, 2017; S.-C. Park, Park, & Hahm, 2011).
This research was designed to evaluate various plant species in order to identify novel antimicrobial peptides through the extraction of protein and peptide fractions. The antimicrobial activity of these extracts was tested against a range of standard bacterial strains, and the peptide nature of the isolated substances was confirmed following their separation and subsequent activity testing. This study builds upon previous investigations that have highlighted the potential of peptides derived from plants as a source of new antibiotics (da Silva et al., 2012; Gully et al., 2019; Lage et al., 2018; Ljoljić Bilić et al., 2022).
During the study, twenty-eight plant seeds, leaves, or flowers were collected from local markets. A phosphate buffer was used to extract water-soluble proteins and peptides, followed by purification through TLC chromatography (Melnikova, Mineev, Finkina, Arseniev, & Ovchinnikova, 2016; Osborn et al., 1995; Taveira et al., 2014). The total protein concentration of the extracts was measured using a NanoDrop instrument, which has been documented in various studies (Jafari, Khavari Nejad, Vaziri, and Siadat 2017; Najib 2017; Desjardins and Conklin 2010).
### a) Antibacterial Activity of Plant Protein Extracts Against Several Standard Bacteria
As part of this study, Ricinus communis seeds protein extract has been studied for its activity against several Gram-positive and Gram-negative standard bacteria. The findings of this study indicate that the protein extract from R. communis demonstrates antibacterial activity against S. aureus and S. epidermidis. The observed antibacterial properties of the protein extract may be linked to the presence of lectins, such as ricin, which are recognized for their antibacterial effects (Al-Mamun et al., 2016), as well as the presence of antimicrobial peptides (Al-Mamun et al., 2016; Boldbaatar, Gunasekera, El-Seedi, & Goransson, 2015).
Numerous studies have evaluated the antimicrobial properties of $R$ communis seed protein extract against various bacteria. For example, research conducted by Al-Mamun et al. (2016) assessed the antibacterial efficacy of the protein extract against $E$ coli, $P$ aeruginosa, and S. aureus through agar well diffusion and MIC assays, yielding MIC values of 250, 125, and $62.5\mu \mathrm{g / ml}$, respectively. Additionally, a study by Afzal, Bakhsh, Ahmad, Manzoor, and Liaquat (2011) utilized the disc diffusion method alongside MIC assays to explore the antibacterial effects of $R$ communisseed protein extract against S. aureus, revealing a MIC value of 15.29 mg/ml. Conversely, research by Patil and Bhise (2015) examined the antimicrobial activity of R.communis seed aqueous extract against several bacteria, including S. aureus and P.aeruginosa, using the agar well diffusion method, and found significant antibacterial activity solely against P. aeruginosa. Furthermore, a study by Abd-Ulgadir, Suliman, Zakria, and Hassan (2015) concluded that the seed protein extract exhibited no notable antimicrobial activity against a variety of Gram-negative and Gram-positive bacteria.
The variations noted in the results of different studies could be attributed to the diverse methodologies used in the protein extraction from seeds (Ben Brahim et al. 2022; Djeussi et al. 2013). Additionally, differences in bacterial strains and the specific experimental culture conditions applied may also contribute to these discrepancies (Ben Brahim et al. 2022). Furthermore, the exact mechanism through which the protein extract functions has not been sufficiently clarified (Worbs et al. 2011).
This study demonstrates that the extract derived from Raphanus sativus seeds exhibited antibacterial properties S. aureus, S. epidermidis, E. coli, and A. baumannii, with concentrations ranging from 0.537 to $2.15\mathrm{mg / ml}$. The antibacterial effects observed may be linked to the presence of compounds such as alkaloids, flavonoids, and saponins, which are likely to be extracted using an aqueous cold extraction method (Ahmad, Hasan, Chishti, & Ahmad, 2012).
Two investigations conducted by Ahmad et al. (2012) and Khamees (2017) assessed the antimicrobial properties of $R$. sativus seed extract against various bacterial strains, including S. aureus, K. pneumoniae, P. aeruginosa, and E. coli. The findings from both studies indicated that the $R$. sativus seed extract demonstrated considerable antibacterial efficacy against all bacterial strains examined. Additionally, both studies revealed the presence of various bioactive compounds, such as alkaloids, flavonoids, and saponins, within the extract, which are believed to play a role in its antibacterial effects. In a related study by Jadoun, Yazbak, Rushrush, Rudy, and Azaizeh (2016), the antibacterial activity of $R$. sativus seed extract was tested against several standard bacterial strains, including S. aureus, Escherichia coli, and K. pneumoniae, with minimum inhibitory concentrations ranging from 0.5 to $1\mathrm{mg / mL}$. The study also identified a novel sulfur compound that contributes to the antibacterial activity. Collectively, these studies indicate that $R$. sativus extract holds promise as a natural antibacterial agent, with potential applications in food preservation and various other industries.
A study conducted by Törün, Çoban, Biyik, and Barişik (2017) examined the antimicrobial properties of methanolic extracts from R. sativus against ten distinct pathogenic microorganisms, such as S. aureus, E. coli, and K. pneumoniae. Utilizing the Disc diffusion method, the findings revealed that the extract exhibited antibacterial activity solely against S. aureus.
In the current study, the growth of S. aureus was found to be inhibited by $2.03\mathrm{mg / ml}$ of Vitis labrusca seeds protein extract. Additionally, the extract was also observed to inhibit the growth of S. epidermidis at a concentration of $4.06\mathrm{mg / ml}$. Few studies have investigated the antibacterial properties of V. labrusca seed extracts. Junior et al. (2021) found that the extracts presented some inhibitory effect against E. coli (IAL 2064) and S. aureus.
In this investigation, the findings indicate that the protein extract derived from *Prunus dulcis* seeds exhibits a minimal antibacterial effect against S. aureus. Nevertheless, the extract failed to show any antibacterial properties against the other bacteria that were examined. Conversely, research conducted by Hifza (2018) on the antimicrobial characteristics of fatty acids extracted from P. dulcis seeds revealed a moderate inhibitory effect against three bacterial strains, including S. aureus, B. subtilis, and E.coli, which aligns partially with the results of the current study.
Additional research has explored the antibacterial properties of $P.$ dulcis seed extract, suggesting that the presence of various bioactive compounds, such as phenolic compounds and flavonoids, may play a role in their antibacterial efficacy (Dhingra, Kar, Sharma, & Bhasin, 2017; Shelly, Shikha, & Narayan, 2015). Consequently, while the protein extract from $P.$ dulcis seeds appears to have limited antibacterial effects against S. aureus, it is plausible that other compounds within the seeds possess more potent antibacterial properties, warranting further investigation to assess their potential applications.
The findings of this research indicate that Hibiscus sabdariffa effectively inhibited the growth of $P$. aeruginosa. Numerous studies have investigated the antibacterial properties of $H$. sabdariffa calyces extract. In 2011, a study found that the methanolic extract showed antibacterial activity against various bacteria (Elmanama et al., 2011). In 2022, Khalil et al. found that the $H$. sabdariffa extract had bactericidal effects against multidrug-resistant $P$. aeruginosa from burn wound exudates. Al-Hashimi's 2012 study found significant inhibition zones for $E$. coli and S. aureus. Vargas-Sanchez et al.'s 2018 study found moderate to high antibacterial activity against various Gram-positive and Gram-negative bacteria.
### b) Thin-layer chromatography (TLC) and TLC agar overlay bioautography for plant protein extracts
During the course of this research, the use of TLC chromatography with agar overlay bioautography to investigate Ricinus communis produced noteworthy findings, demonstrating two separate zones of inhibition on the agar plate against S. aureus and S. epidermidis. One zone corresponded to a fraction with an Rf value of
0.06, while the other was linked to a fraction with an Rf value of 0.73. These zones of inhibition may be attributed to the presence of lectins, such as ricin, known for their antibacterial properties, as well as antimicrobial peptides, as indicated by Al-Mamun et al. (2016). Further investigations are necessary to isolate and characterize the chemical nature of each fraction. In a separate study conducted by Lekganyane (2015), TLC chromatography was employed to separate phytochemical compounds from various leaf extracts, including that of $R.$ communis. The resulting fractions were subjected to bioautography to evaluate their antimicrobial efficacy against different bacterial strains. The study revealed that a fraction with an Rf value of 0.77 demonstrated significant antimicrobial activity against S. aureus.
The antibacterial activity of V. labrusca seed extract was assessed using TLC plates and a TLC agar overlay bioautography assay. The results showed two stained fractions but no antibacterial effects against S. aureus and S. epidermidis. A zone of inhibition was noted near the sample application line, suggesting that non-proteinaceous compounds may be responsible for the antibacterial effects. Previous studies identified bioactive polyphenols like anthocyanins, flavonoids, and resveratrol in V. labrusca, which are known for their antioxidant, anti-inflammatory, and antimicrobial properties (Xia, Deng, Guo, & Li, 2010; Yu & Ahmedna, 2013). These compounds likely contribute to the antibacterial activity observed in the extract.
In the context of this study, the thin-layer chromatography (TLC) analysis of Raphanus sativus seed extract revealed two overlapping bands, neither of which demonstrated any antibacterial properties. Nevertheless, a notable zone of inhibition was detected adjacent to the sample application line on the TLC plate when tested against S. aureus, S. epidermidis, E. coli, and A. baumannii. The findings suggest that the antibacterial effects of $R.$ sativus extract are unlikely to be linked to the proteins or peptides present, as evidenced by the lack of antibacterial activity in the two protein bands observed. Instead, the inhibition zone implies that other constituents within the extract may be responsible for the antibacterial effects. Research has pointed to sulfur compounds, particularly isothiocyanates, as significant contributors to the antibacterial properties of $R.$ sativus extract, with sulfuraphane being a prominent isothiocyanate known for its efficacy against various bacterial strains (Gutiérrez & Perez, 2004; Jadoun et al., 2016; Lim, Han, & Kim, 2016). The presence of these sulfur compounds may elucidate the antibacterial activity observed in $R.$ sativus extract, underscoring its potential as a natural source of antimicrobial agents In this research, the antibacterial properties of the extract derived from Prunus dulcis seeds were assessed. Interestingly, although the agar well diffusion
method indicated minimal antibacterial effects against S. aureus, the agar overlay bioautography did not reveal any significant antibacterial activity from the extract. This discrepancy may suggest that the concentration of antibacterial compounds present in the extract was insufficient to be detected by the agar overlay bioautography technique (Bieleski & Turner, 1966).
In the course of this investigation, the protein extract from the calyces of Hibiscussabdariffa was subjected to thin-layer chromatography (TLC), which successfully identified three distinct stained bands on the chromatographic plate. Subsequent evaluation of the $H.$ sabdariffa extract against $P.$ aeruginosa through agar overlay bioautography demonstrated the presence of several large zones of inhibition, a stark contrast to the smaller zone noted in the agar well diffusion assay. This discrepancy in findings may be attributed to the potential antagonistic interactions among the extract's components during the agar well diffusion assay; however, following separation via TLC, these compounds likely ceased to interact, thereby allowing for individual antimicrobial activity and resulting in the observed larger inhibition zones in the agar overlay bioautography. The phenomenon of antagonism among constituents within a single plant extract has also been documented in research conducted by Vaou et al. (2022) regarding artemisinins, which are antimalarial compounds derived from Artemisia annua. Additionally, another plausible reason for the variation in outcomes between the agar well diffusion assay and the agar overlay bioautography could be that the TLC process eliminated certain interfering substances that were present in the original extract, as noted by Bieleski and Turner (1966).
The identification of several zones of inhibition against $P$ aeruginosa indicates that the plant extract is likely composed of various bioactive compounds with antibacterial properties. Notably, only one of these inhibition zones was linked to the protein fraction, while the remaining two zones were located close to the sample application line and the edge of the TLC plate. This observation implies that a limited aspect of the antibacterial effect may be due to the presence of antimicrobial peptides. Furthermore, there is a lack of prior research that has successfully isolated or evaluated antimicrobial peptides derived from the extract of $H$ sabdariffa calyx.
In the present work, the antibacterial properties of several plant extracts, including Vitis labrusca, Ricinus communis, Raphanus sativus, Prunus dulcis, and Hibiscus sabdariffa, were examined through various methodologies such as agar well diffusion assay, micro broth dilution, bioautography, and thin-layer chromatography (TLC). The findings indicated that the antibacterial effects of these extracts could not be exclusively linked to the presence of proteins or peptides; rather, other phytochemicals, including polyphenols, flavonoids, tannins, and alkaloids, may play a significant role in their efficacy. The specific active compounds were not identified, which points to the necessity for additional investigations employing advanced techniques like mass spectrometry and nuclear magnetic resonance (NMR) spectroscopy. The study also highlighted the intricate nature of plant extracts and the difficulties associated with pinpointing their active constituents. Future research endeavors may focus on the isolation and characterization of individual compounds, as well as exploring their mechanisms of action against bacterial strains. Overall, this investigation offers valuable insights into the potential of plant extracts as antibacterial agents and emphasizes the critical need for continued research in this field.
Disclosure statement: The authors declare no conflict of interest.
Generating HTML Viewer...
References
59 Cites in Article
K Abd-Ulgadir,S Suliman,I Zakria,N Hassan (2015). Antimicrobial potential of methanolic extracts of Hibiscus sabdariffa and Ricinus communis.
I Abed (2015). Antibacterial Effect of Flavonoids Extracted from Seeds of Silybum marianum against Common Pathogenic Bacteria.
E Abou-Elkhair,H Fadda,U Abu-Mohsen (2010). Antibacterial activity and Phytochemical analysis of some medicinal plants from Gaza Strip-Palestine.
M Afzal,A Bakhsh,M Ahmad,I Manzoor,S Liaquat (2011). COMPARISON OF THE ANTIMICROBIAL ACTIVITY OF SEED PROTEIN EXTRACTSFROM SIX MEDICINAL PLANTS AGAINST STAPHYLOCOCCUS AUREUS 6736152.
F Ahmad,I Hasan,D Chishti,H Ahmad (2012). Raphanus Sativus Linn., Crude Extract Compound 53123‐88‐9.
Kabir Akinyemi,Olukayode Oladapo,Chidi Okwara,Christopher Ibe,Kehinde Fasure (2005). Screening of crude extracts of six medicinal plants used in South-West Nigerian unorthodox medicine for anti-methicillin resistant Staphylococcus aureus activity.
A Al-Hashimi (2012). Five Plants with Promising Antimicrobial Peptides Global Journal of Medical Research.
M Al-Mamun,Zerin Akter,Md Uddin,K Ferdaus,K Hoque,Z Ferdousi,M Reza (2016). Characterization and evaluation of antibacterial and antiproliferative activities of crude protein extracts isolated from the seed of Ricinus communis in Bangladesh.
A Aliahmadi,R Roghanian,G Emtiazi,A Ghassempour (2011). Agar-overlay assay; a useful and cost benefit method for detection of antibacterial peptides in plant seeds.
Luís Almeida,Caio Oliveira,Mayara Rodrigues,Simone Neto,Ana Boleti,Gabriel Taveira,Érica Mello,Valdirene Gomes,Edson Santos,Edson Crusca,Octávio Franco,Marlon Cardoso,Maria Macedo (2020). Adepamycin: design, synthesis and biological properties of a new peptide with antimicrobial properties.
Uttpal Anand,M Carpena,Monika Kowalska-Góralska,P Garcia-Perez,Kumari Sunita,Elza Bontempi,Abhijit Dey,Miguel Prieto,Jarosław Proćków,Jesus Simal-Gandara (2022). Safer plant-based nanoparticles for combating antibiotic resistance in bacteria: A comprehensive review on its potential applications, recent advances, and future perspective.
Atanas Atanasov,Birgit Waltenberger,Eva-Maria Pferschy-Wenzig,Thomas Linder,Christoph Wawrosch,Pavel Uhrin,Veronika Temml,Limei Wang,Stefan Schwaiger,Elke Heiss,Judith Rollinger,Daniela Schuster,Johannes Breuss,Valery Bochkov,Marko Mihovilovic,Brigitte Kopp,Rudolf Bauer,Verena Dirsch,Hermann Stuppner (2015). Discovery and resupply of pharmacologically active plant-derived natural products: A review.
Sylvia Baltzer,Melissa Brown (2011). Antimicrobial Peptides – Promising Alternatives to Conventional Antibiotics.
Anna Barashkova,Eugene Rogozhin (2020). Isolation of antimicrobial peptides from different plant sources: Does a general extraction method exist?.
Raoua Ben Brahim,Hasna Ellouzi,Khaoula Fouzai,Nedra Asses,Mohammed Neffati,Jean Sabatier,Philippe Bulet,Imed Regaya (2022). Optimized Chemical Extraction Methods of Antimicrobial Peptides from Roots and Leaves of Extremophilic Plants: Anthyllis sericea and Astragalus armatus Collected from the Tunisian Desert.
R Bieleski,N Turner (1966). Separation and estimation of amino acids in crude plant extracts by thin-layer electrophoresis and chromatography.
Delgerbat Boldbaatar,Sunithi Gunasekera,Hesham El-Seedi,Ulf Göransson (2015). Synthesis, Structural Characterization, and Bioactivity of the Stable Peptide RCB-1 from <i>Ricinus communis</i>.
Marlon Cardoso,Beatriz Meneguetti,Bruna Costa,Danieli Buccini,Karen Oshiro,Sergio Preza,Cristiano Carvalho,Ludovico Migliolo,Octávio Franco (2019). Non-Lytic Antibacterial Peptides That Translocate Through Bacterial Membranes to Act on Intracellular Targets.
Pooi Chung,Ramona Khanum (2017). Antimicrobial peptides as potential anti-biofilm agents against multidrug-resistant bacteria.
Marjorie Cowan (1999). Plant Products as Antimicrobial Agents.
B Da Silva,V De Freitas,L Nascimento-Neto,V Carneiro,F Arruda,A De Aguiar,. Teixeira,E (2012). Antimicrobial peptide control of pathogenic microorganisms of the oral cavity: a review of the literature.
Philippe Desjardins,Deborah Conklin (2010). NanoDrop Microvolume Quantitation of Nucleic Acids.
N Dhingra,A Kar,R Sharma,S Bhasin (2017). In-vitro antioxidative potential of different fractions from Prunus dulcis seeds: Vis a vis antiproliferative and antibacterial activities of active compounds.
D Djeussi,J Noumedem,J Seukep,A Fankam,I Voukeng,S Tankeo Unknown Title.
V Kuete (2013). Antibacterial activities of selected edible plants extracts against multidrug-resistant Gram-negative bacteria.
A Elmanama,A Alyazji,N Abu-Gheneima (2011). Antibacterial, antifungal and synergistic effect of Lawsonia inermis, Punica granatum and Hibiscus sabdariffa.
C Ghosh,P Sarkar,R Issa,J Haldar (2019). Alternatives to conventional antibiotics in the era of antimicrobial resistance.
Kay Gully,Sandra Pelletier,Marie-Charlotte Guillou,Marina Ferrand,Sophie Aligon,Igor Pokotylo,Adrien Perrin,Emilie Vergne,Mathilde Fagard,Eric Ruelland,Philippe Grappin,Etienne Bucher,Jean-Pierre Renou,Sébastien Aubourg (2019). The SCOOP12 peptide regulates defense response and root elongation in <i>Arabidopsis thaliana</i>.
Rosa Gutiérrez,Rosalinda Perez (2004). Raphanus sativus (Radish): Their Chemistry and Biology.
Radosław Gwarda,Aleksandra Tomczyszyn,Aleksandra Misicka,Tadeusz Dzido (2013). Staining of some synthetic oligopeptides using ninhydrin solution.
Behnoush Jafari,Ramazan Khavari Nejad,Farzam Vaziri,Seyed Siadat (2017). Evaluation of the effects of extracellular vesicles derived from Faecalibacterium prausnitzii on lung cancer cell line.
Maciej Jaskiewicz,Malgorzata Orlowska,Gabriela Olizarowicz,Dorian Migon,Daria Grzywacz,Wojciech Kamysz (2016). Rapid Screening of Antimicrobial Synthetic Peptides.
Najwan Jubair,Mogana Rajagopal,Sasikala Chinnappan,Norhayati Abdullah,Ayesha Fatima (2021). Review on the Antibacterial Mechanism of Plant-Derived Compounds against Multidrug-Resistant Bacteria (MDR).
Tufy Junior,Cristiane De Moura,Mariana Do Carmo,Luciana Azevedo,Luis Esmerino,Rosangela Tardivo,Petri Kilpeläinen,Daniel Granato (2021). Chemical Composition, Antioxidant, Antimicrobial and Cytotoxic/Cytoprotective Activity of Non-Polar Extracts of Grape (Vitis labrusca cv. Bordeaux) and Blackberry (Rubus fruticosus) Seeds.
Wojciech Kamysz,Marcin Okrój,Jerzy Łukasiak (2003). Novel properties of antimicrobial peptides..
Rania Khalil,Galal Yahya,Walied Abdo,Ghada El-Tanbouly,Dina Johar,Mahmoud Abdel-Halim,Hanan Eissa,Calin Magheru,Sameh Saber,Simona Cavalu (2022). Emerging Approach for the Application of Hibiscus sabdariffa Extract Ointment in the Superficial Burn Care.
A Khamees (2017). Phytochemical and pharmacological analysis for seeds of two varieties of Iraqi Raphanus sativus.
E Klein,T Van Boeckel,E Martinez,S Pant,S Gandra,S Levin,. Laxminarayan,R Hancock (2001). Global increase and geographic convergence in antibiotic consumption between 2000 and 2015.
Evan Haney,Robert Hancock (2013). Peptide design for antimicrobial and immunomodulatory applications.
A Hifza (2018). Extraction and Bioanalysis of Ultrasonic Assisted Pakistani Cultivar Prunus Dulcis Seed: An Optimization Study.
Jeries Jadoun,Ahmad Yazbak,Salwa Rushrush,Amira Rudy,Hassan Azaizeh (2016). Identification of a New Antibacterial Sulfur Compound from <i>Raphanus sativus</i> Seeds.
R Osborn,G De Samblanx,K Thevissen,I Goderis,S Torrekens,F Van Leuven Unknown Title.
W Broekaert (1995). Isolation and characterisation of plant defensins from seeds of Asteraceae, Fabaceae, Hippocastanaceae and Saxifragaceae.
Seong-Cheol Park,Yoonkyung Park,Kyung-Soo Hahm (2011). The Role of Antimicrobial Peptides in Preventing Multidrug-Resistant Bacterial Infections and Biofilm Formation.
Sakhalkar Sachin,Joshi Prachi (2015). Study of Antioxidant Activity in Vitro by Aqueous, Ethanolic and Ether Extract of Hibiscus Rosa-Sinensis Linn.
M Sharma,R Abid,M Sajgotra (2017). Phytochemical screening and thin layer chromatography of Ficus carica leaves extract.
A Shelly,M Shikha,S Narayan (2015). FUNCTIONAL PROPERTIES OF YOGURT PREPARED USING A BLEND OF ALMOND MILK (Prunus dulcis) AND PEANUT MILK (Arachis hypogaea L.).
James Tam,Shujing Wang,Ka Wong,Wei Tan (2015). Antimicrobial Peptides from Plants.
S.-S Tang,Z Prodhan,S Biswas,C.-F Le,S Sekaran (2018). Antimicrobial peptides from different plant sources: Isolation, characterisation, and purification.
Gabriel Taveira,Luciana Mathias,Olney Da Motta,Olga Machado,Rosana Rodrigues,André Carvalho,André Teixeira‐ferreira,Jonas Perales,Ilka Vasconcelos,Valdirene Gomes (2014). Thionin‐like peptides from <i>Capsicum annuum</i> fruits with high activity against human pathogenic bacteria and yeasts.
Bahadir Törün,Esin Çoban,Esra Barişik (2017). Antimicrobial Activity of Echinophora tenuifolia L. and Raphanus sativus L. Extracts.
L Tzong-Hsien,N Kristopher,A Marie-Isabel (2016). Antimicrobial Peptide Structure and Mechanism of Action: A Focus on the Role of Membrane Structure.
Jill Winkler-Moser (2022). Variations in Phytochemicals in DDGS Oil from 30 Ethanol Plants.
D Valle,Jr,J Puzon,E Cabrera,W Rivera (2016). Thin Layer Chromatography-Bioautography and Gas Chromatography-Mass Spectrometry of Antimicrobial Leaf Extracts from Philippine Piper betle L. against Multidrug-Resistant Bacteria.
Natalia Vaou,Elisavet Stavropoulou,Chrysoula Voidarou,Zacharias Tsakris,Georgios Rozos,Christina Tsigalou,Eugenia Bezirtzoglou (2022). Interactions between Medical Plant-Derived Bioactive Compounds: Focus on Antimicrobial Combination Effects.
Who (2017). Sibson, Prof. Robin, (4 May 1944–19 March 2017), Chief Executive, Higher Education Statistics Agency, 2001–09 (Board Member, 1996–2000).
S Worbs,K Köhler,D Pauly,M.-A Avondet,M Schaer,M Dorner,B Dorner (2011). Ricinus communis intoxications in human and veterinary medicine-a summary of real cases.
E.-Q Xia,G.-F Deng,Y.-J Guo,H.-B Li (2010). Biological activities of polyphenols from grapes.
Jianmei Yu,Mohamed Ahmedna (2013). Functional components of grape pomace: their composition, biological properties and potential applications.
No ethics committee approval was required for this article type.
Data Availability
Not applicable for this article.
How to Cite This Article
Hend N. Essawaf. 2026. \u201cFive Plants with Promising Antimicrobial Peptides\u201d. Global Journal of Medical Research - C: Microbiology & Pathology GJMR-C Volume 25 (GJMR Volume 25 Issue C1): .
Explore published articles in an immersive Augmented Reality environment. Our platform converts research papers into interactive 3D books, allowing readers to view and interact with content using AR and VR compatible devices.
Your published article is automatically converted into a realistic 3D book. Flip through pages and read research papers in a more engaging and interactive format.
Antimicrobial peptides (AMPs) are bioactive molecules known for their strong antimicrobial properties against a variety of microorganisms. The increasing prevalence of antibiotic-resistant infections, coupled with a scarcity of newly developed antibiotics, has intensified the search for novel sources of AMPs. Plant-derived AMPs present several benefits compared to conventional antibiotics. Consequently, this research focused on screening various plant species to identify new antimicrobial peptides. Water-soluble proteins were extracted from the dried plant materials using an aqueous extraction buffer. The resulting extracts were evaluated for antibacterial activity, with the minimum inhibitory concentration (MIC) being determined, followed by separation through thin layer chromatography and subsequent agar overlay bioautography. Protein extracts from five different plants exhibited antibacterial properties against both Gram-positive and Gram-negative bacteria, with inhibition zones measuring between 11 and 22 mm.
Our website is actively being updated, and changes may occur frequently. Please clear your browser cache if needed. For feedback or error reporting, please email [email protected]
Thank you for connecting with us. We will respond to you shortly.