Infection of SARS-COV2 and its variants causes wide range morbidity and mortality in recent years. Identification of epitope-based mechanism of viral infection with progressive fatality and antiviral immunotherapy are two major goals to address population-bias immune response. We selected peptides from SARS-COV2 Spike (6VXX_A), Delta (B.1.617.2), Omicron (B.1.1.529) proteins. These peptides contain epitopes which are identified as low rank good fit immunogenic as recognized more by HLA-DRB1*15:01, than HLA-DRB1*07:01 and HLA-DRB1*03:01. We also found the selected epitopes specifically form interactive complex with mucosa-associated invariant T cell. The Molecular Docking and Molecular Dynamics experiments demonstrated amino acid sequence-specific interaction between close atoms from epitopes and MAIT-TCR. We used virus unrelated microbial peptide antigen85 as control.
## I. INTRODUCTION
Severe Acute Respiratory Syndrome Corona Virus-2 (SARS-COV2) infection caused pandemic (COVID-19) in recent years 2019-2021 with high mortality rate (1-4). The pre-existing pulmonary conditions worsen the clinical outcome during infection. Recent emergence of Delta and Omicron variants with advent of subvariants clearly indicate a range of fast transforming viruses appear in diverse human population (5-7). The genetic predisposition for intrinsic susceptibility of human populations towards SARS-COV2 and variants Delta, Omicron is still unclear. Also, the specific role of T helper cells, its receptors and T cell repertoires in viral antigen recognition for antiviral immune response is not known yet. Several reports suggested involvement of HLA responses in SARS-COV2 infection and after vaccination (8-10). Though we still have unresolved questions to address: (1) HLA allele specificity for antigen presentation and population bias expression patterns; (2) T helper cell expansion and TCR specificity towards viral antigens. The reports from different laboratories suggest HLA-DR deficiency or low expression in monocytes is associated with immunosuppression and severity in SARS-COV2 infected patients (11-14). In particular, specificity of HLA-DRB1 alleles to recognize Delta and Omicron variants of SARS-COV2 and T cell receptor specificity towards epitopes is unclear yet but critical for evaluating hosts' protective immune response.
We identified immunogenic epitopes from N-terminal first nine hundred-amino acid sequence of SARS-COV2 (COVID 19), Delta variant (B.1.617.2) and Omicron Variant (B.1.1.529) Spike proteins. We found HLA-DRB1 allele specificity for these selected Spike epitopes is determinant for recognition by mucosa-associated invariant T cell receptor in human population.
## II. MATERIALS AND METHODS
a) SARS-COV2 and its variant protein sequences
- The N-terminal first 900 amino acid Spike protein sequence is selected from NCBI protein databases (SARS-COV2: 6VXX_A, Delta variant (B. 1.617.2): 7ORB_R, Omicron (B.1.1.529): 7Q09_A. The protein sequence was processed by NCBI protein blast (pblast) software engine to determine 60 amino acid peptide strings. The randomly selected 25-30 Spike peptide strings containing epitopes were processed for protein database (PDB) file by using Avogadro (USA). The Omicron Spike epitope PDB files: OM42, OM60, M312, M370 and SARS-COV2 (6VXX_A)-Delta variant (7ORB_R) overlapping CD4 immunogenic epitope -PDB file DLT1, Delta variant (7ORB_R) epitope DLT2 (Table 1) are used to determine close alignment and binding efficiency with mucosa associated invariant T cell receptor Vα7.2Vβ7.2 (MAIT-TCR).
b) Epitope sequence identification from Spike protein of SAR-COV2, Delta, Omicron Variants
The CD4- recognizing epitope sequences were identified from N terminal nine hundred amino acid sequences of virus Spike envelope protein. The amino acid sequences were analyzed for their immunogenicity by Immune Epitope Database (IEDB) software engine
(NIAID, USA). The IEDB Tools were programmed to analyze 7 allele response to virus Spike proteins following methods described by Dhanda et al. (15). The HLA-DRB1*03:01, HLA-DRB1*07:01 and HLA-DRB1*15:01 responsive epitopes were selected for determining specific binding affinity with MAIT- TCR.
### c) Molecular Docking and Molecular dynamics
The selected peptide epitopes from N terminal first nine hundred amino acid sequence are processed by Avogadro software (USA) for construction of protein database (PDB). These PDB sequences are allowed to run (100 nanosecond) for close alignment with mucosa associated invariant T cell receptor (PDB: 4L9L). The specifications include recognition of alpha helical structure for interaction, speed 30 frames per second with Ray trace. The epitopes are placed as mobile and the TCR is placed as target. The molecular dynamics experiments ran for 10 cycles with cut off value 2 Angstrom unit. After completion of the run, the frame per nanosecond was calculated under Executive Root Mean Square Deviation (RMSD) value ranging from 5.2-6.2 Angstrom unit (RMSD: 2 Angstrom indicates good homology between interacting protein sequences thus closer to bind together). The aligned interacting peptide epitope with TCR protein sequence is recorded by Molecular Dynamics method by using Schrodinger PyMol software (USA). The Molecular Docking of Spike epitopes on TCR is determined in the same experiment at different time points.
### d) Statistics
ANOVA is used to determine significance $(p<0.05$ at least) level of interaction profiles (Frame/ nanosec) between selected virus epitope and with respect to unrelated virus peptides. Mycobacterium avium antigen85 peptide was used as control to determine significant binding of virus epitopes with MAIT-TCR selective amino acid sequence.
## III. RESULTS
T cell specificity of SARS-COV2 Spike peptides is critical for prolongant antiviral responses leading towards development of memory. However, it is unclear whether localized TCR - immune response in mucosal layers have specificity towards SARS-COV2 -Spike epitopes? It is also unclear whether Cytotoxic T cells (CD8), natural killer T cells (NKT) and other localized invariant T cells have specific response towards HLA-DRB1 allele specific immunogenic Spike epitopes.
In order to address the questions, we first screened the viral epitopes derived from N (NH2)-terminal first 900 amino acid sequence of SARS-COV2 and its variant Delta, Omicron (Described in Methods section). Then, we selected each of these epitopes and analyzed their immunogenicity with respect to binding with HLA-DRB1*03:01, HLA-DRB1*07:01 and HLA-
DRB1*15:01. These HLA-DRB1 allele recognized viral epitopes are checked for their efficacy for CD4 and TCR recognition. Here, we presented recognition ability of mucosa-associated invariant T cell receptor (MAIT-TCR) to the selected epitopes derived from Spike envelop protein sequences of SARS-COV2, Delta and Omicron variants.
### a) Recognition of N-terminal Spike epitopes of SARS-COV2 and Delta variant by HLA-DRB1 alleles
The results presented in the Figure 1 show differential responses of screened N-terminal Spike epitopes of SARS-COV2 (ID 6VXX_A) (Fig.1A) and Delta variant (7ORB_R) (Fig.1B) towards HLA-DRB1 alleles 03:01, 07:01 and 15:01. The bar diagrams with epitope database analysis tool (IEDB) percentile ranks demonstrate a base line of 20 percentile; below which the epitopes are low immunogenic and good fit to HLA-DRB1 alleles. These epitopes have ability to induce moderate to low CD4 responses. The results showed overlapping recognition of epitope LYNSASFST (derived from 6VXX_A and 7ORB_R) by HLA-DRB1*03:01 and HLA-DRB1*15:01 (< 20 percentile rank) for SARS-COV2 and Delta variant. HLA-DRB1*07:01 recognition of the same epitope showed marginally over than 20 percentile rank. The epitope has lower rank (45.38232) when it is recognized by HLA-DRB1*15:01 and HLA-DRB1*03:01. The epitope YFKIYSKHT (rank: 42.8908) derived from SARS-COV2 Spike protein (ID 6VXX_A) is recognized by HLA-DRB1*15:01 and HLA-DRB1*07:01 while another epitope peptide VSLLSVLLM/ LVSLLSVLL (rank: 50.16648/ 52.20664) is recognized by all three tested HLA- DRB1*03:01, HLA-DRB1*07:01 and HLA-DRB1*15:01alleles. The specific recognition by HLA-DRB1*07:01 and HLA-DRB1*15:01 allele is detected for Delta variant epitope FASYAWNR (rank: 43.9931), YAWNRKRIS (rank: 40.04248), YRLFRKSNL (34.09812). These HLA-DRB1 selected Spike epitopes from SARS-COV2 and Delta variants are immunogenic to CD4 T cells in human populations.
### b) Recognition of N-terminal Spike epitopes of Omicron variant by HLADRB1 alleles
The recently detected Omicron variant (B.1.1.529) in post pandemic period human population raise another question on protective antiviral responses of existing vaccines and drugs. The presence of several mutations in the virus RNA genome with possible quick adaptation ability in human population bring the variants as highly infectious and transmissible subtypes of SARS- COV2. To determine epitope recognition patterns by HLA-DRB1 alleles, the N-terminal amino acid sequences of Omicron variant (B.1.1.529) Spike protein was selected from protein database, Sequence ID 7Q07_A (NCBI) and screened for CD4 immunogenicity by using IEDB Tools software as mentioned in the Methods section above. The first one thousand amino acid containing sequence was processed for constructing protein database (PDB) files. The Omicron PDB files OM42 (amino acid 1-42); OM60 (amino acid 43-60); M312 (amino acid 312-342), M370 (amino acid 343-370) are constructed similarly as mentioned above. The Table 2 demonstrates overlapping 15-amino acid Omicron peptide sequences (highlighted part of the sequences). The first three epitope peptides in the Table 2 show the lowest rank (5.11, 9.77 and 21) indicating good fit recognition by HLA-DRB1*15:01 as compared with HLA-DRB1*03:01 and HLA-DRB1*07:01. The peptide epitopes in the Table 2 demonstrate moderate to higher HLA-DRB1 recognition patterns not only for 15:01 but 03:01 and 07:01 alleles, thus range poor to non-immunogenic to CD4.
### c) Mucosa associated invariant T cell receptor (MAIT-TCR) binding with Spike epitopes
The specificity of virus epitope induced activation of human immune system is determined by demonstration of sequence specific recognition of virus epitopes with T cell receptor. In the aspect, we wanted to determine the interaction and close binding profiles of TCR and our selected CD4 immunogenic virus epitopes. We found; selected epitopes are recognized by CD4 to a variable extent generating over all low to moderate CD4 response (with respect to HLA-DR recognition pattern). The mucosal associated invariant T cells (MAIT) are conserved lineage of CD4-CD8-/CD4-CD8+ T cell subset in human population (16-19). MAIT cells express Valpha7.2/Jalpha33 alpha chain paired with Vbeta2 beta chain. The MAIT-TCR Valpha domain has similarity with type 1 invariant natural killer T (iNKT) cells like CD1 positive cells which respond to different microbial pathogens and autoimmune manifestations (20).
We wanted to determine efficacy of the selected immunogenic SARS-COV2 and variant Spike epitopes (Table1) to form close binding interaction with MAIT-TCR (PDB: 4L9L).
The protein database (PDB) files of virus epitopes (Table 1) are allowed to interact with mucosal associated invariant T cell receptor (MAIT-TCR) by molecular docking and molecular dynamics experiments. Figure 2A shows efficient binding between DLT2 and TCR 4L9L at the amino acid sequence 1 to 9 region (Square box showing specific alignment between epitope sequence DLT2 and TCR 4L9L). The RMSD value: 2.309 (Fig. 2B) indicates close alignment with more sequence homology thereby efficient binding between DLT2 epitope and MAIT- TCR.
The figure 2C demonstrates alignment between interacting Delta epitope DLT1 and MAIT-TCR at amino acid sequence 1 to 9 region. The efficiency of binding between DLT1 epitope and TCR 4L9L is determined by root mean square deviation (RMSD) value: 5.410 Angstrom (Fig. 2D). The RMSD value indicates moderate to less sequence homology between interacting peptides thereby moderate to low binding as compared with RMSD value for DLT2 and TCR interaction.
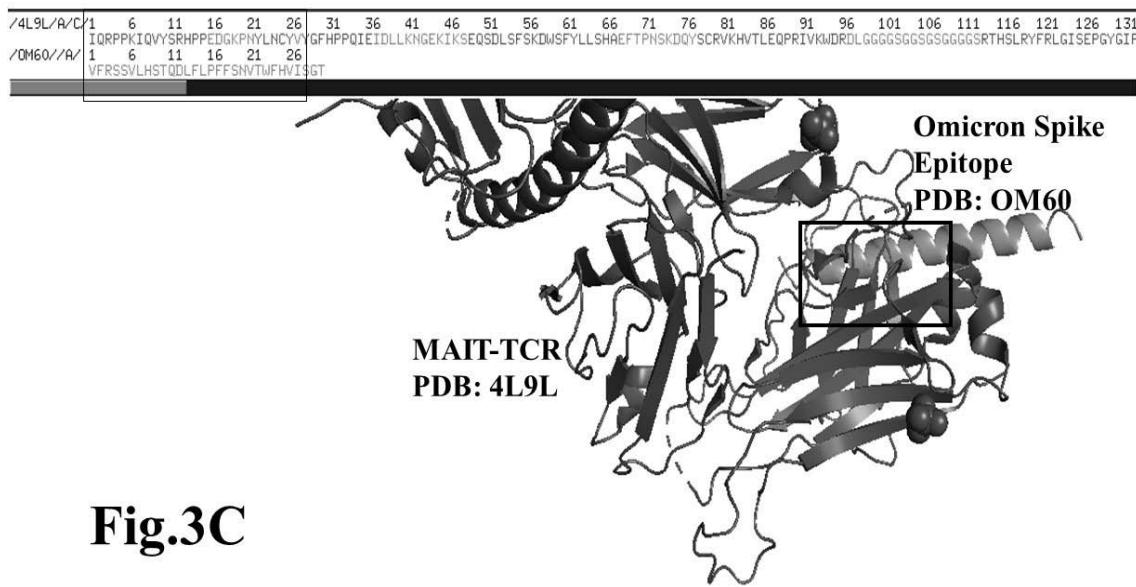
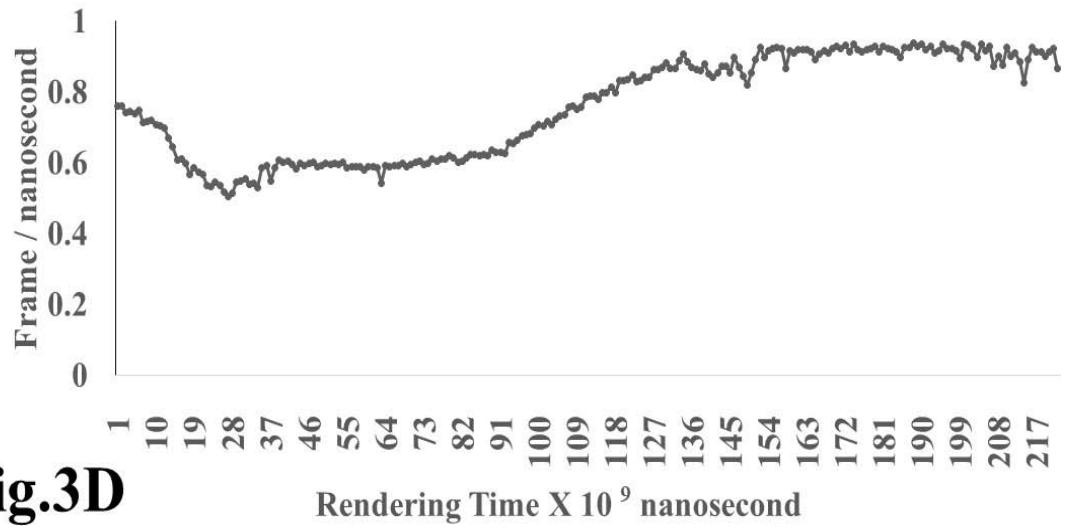
The results presented in figure 3 A demonstrate an aligned binding between Omicron Spike epitope (PDB: OM42) and MAIT- TCR (PDB: 4L9L) sequences. The arrowhead and a square box show the region of binding of OM42 with TCR protein sequence. The digital experimentation using Schrodinger PyMol software identified 1-6 amino acid sequence in TCR binds with OM42 Omicron epitope (the square box indicates aligned amino acid sequences). The 100-nanosecond run with 30 frames/ nanosecond output for 10 cycles demonstrates atom to atom interaction profile between target TCR and mobile OM42 (Fig.3B). The executive root mean square deviation (RMSD) value 6.612 Angstrom shows relative differences between interacting atoms from epitope OM42 and TCR 4L9L. The epitope OM60 binds with MAIT- TCR at 1-26 amino acid region (square box shows the sequence interactions) (Fig.3C). Corresponding molecular dynamics show higher root mean square value (RMSD: 7.282) (Fig.3D). The observation indicates less alignment between interacting atoms of OM60 and TCR 4L9L.
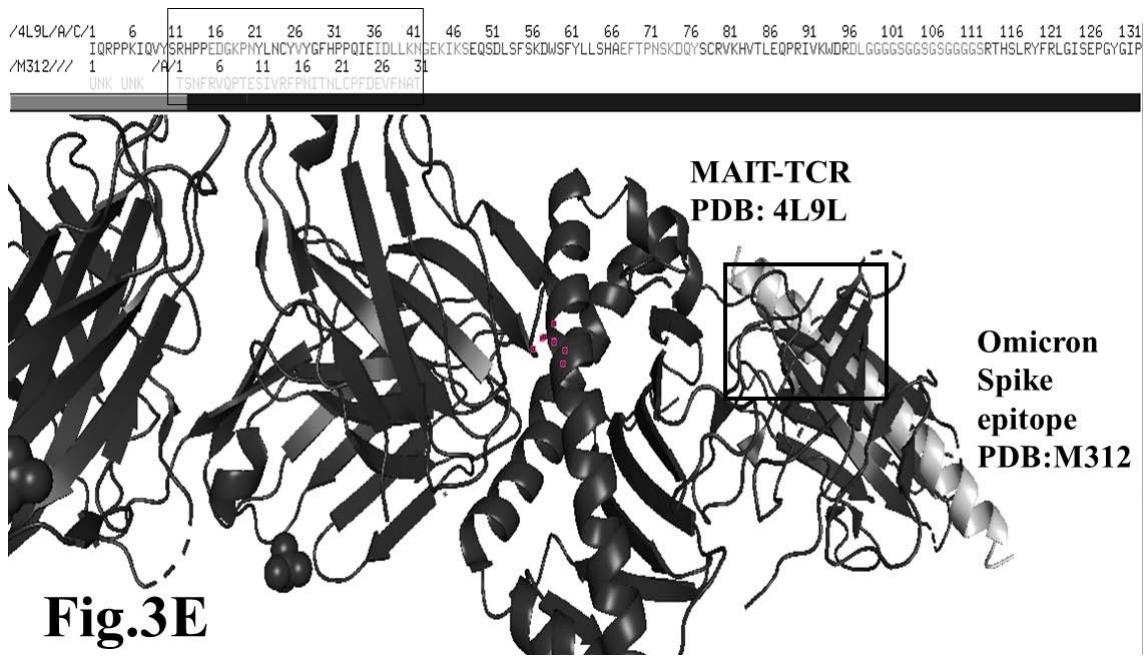
Omicron epitope M312 (amino acid sequence 1-31) is shown to bind with TCR 4L9L (amino acid sequence 11-41) (Fig. 3E). The corresponding molecular dynamics experiments demonstrate root mean square deviation (RMSD) value 6.131 Angstrom which indicate less sequence homology, more deviation between interacting atoms leading toward moderate binding between Omicron epitope M312 and MAIT TCR (Fig.3F). The epitope M370 demonstrates binding of M370 amino acid sequence 1 to 26 with MAIT-TCR 4L9L amino acid sequence 1 to 26 (the box shows interacting sequences) (Fig. 3G). The molecular dynamics experiments show 100 nanoseconds run with 30 frames per nanosecond in which atomic interactions between Omicron epitope M370 and MAIT-TCR exhibit recognizable differences with moderate alignment between two different sequences close enough to bind each other. The root- mean square value RMSD: 6.364 Angstrom indicate difference between the atomic interactions with rendering time (Fig. 3H).
The specificity of viral epitope binding with MAIT-TCR (4L9L) is determined by deletion peptide sequences in which sequential three amino acid deletion was performed to construct new set of peptides. These deletion peptide constructs were processed for PDB files and allowed to interact with MAIT -TCR. The root- mean square deviation (RMSD) (5 to $\leq 2$ Angstrom) values demonstrate specific binding of virus epitope with TCR (data not shown).
In order to find out specificity of binding between virus Spike envelop protein epitopes and MAIT-TCR in the molecular dynamics experiments, we use single factor ANOVA to determine significant differences between number of frame per nanosecond values derived from Omicron Spike epitopes OM42, OM60, M312, M370 and SAR-COV2, Delta variant epitopes DLT1, DLT2 from control peptide: antigen85 (Mycobacterium avium soluble antigen, unrelated to virus protein). The results demonstrate significance level at $p < 0.05$ between control number of frame per nanosecond and that of all Omicron epitopes interacting with MAIT-TCR in molecular dynamics experiments. The frame per second value of SAR-COV2 and Delta epitope also demonstrate significant differences ( $p < 0.05$ ) with respect to control molecular dynamics with MAIT-TCR.
The observations suggest such a marked difference in receptor-ligand interaction patterns between SAR-COV2, Delta and Omicron peptide epitopes with MAIT-TCR as compared with unrelated control peptide antigen85 is due to difference in mean oscillation pattern between interacting atoms of receptor and ligands close to each other. The lower the RMSD value the closer the interacting atoms are so probability of bond formation increases.
## IV. DISCUSSION
SARS-COV2 and its variants Delta and recent emergent Omicron viruses cause severe to mild respiratory tract infection in worldwide human population. The available reports from 2019 till ongoing 2022, we found severity of infection and mortality during pandemic to post pandemic period depends largely on uncontrolled heightened immune response leading inflammation and tissue damage (21, 22). However, recognition patterns of SARS-COV2 Spike protein epitopes to T cell repertoire and long-term memory immune response is still unclear. We consider the (a) antigenicity of Spike epitopes on its recognition to HLA-DRB1 alleles and (b) optimum T cell receptor response to HLA-DRB1 allele(s)-recognized epitopes are two requirements to activate T helper cells for clonal expansion and development of memory cell pool. So far, we do not see distinct memory cell pool and expansion of T helper cell repertoire during secondary infection in human population. The experimental vaccines show 6 months' time period of moderate protection. Here the question is whether the Spike peptide epitopes have ability to activate T cytotoxic (CD8) cells and Natural Killer cells more than T helper cells? We tested T helper cell (CD4) immunogenicity and generalized T cell receptor (TCR) responses to first thousand N - terminal Spike epitopes screened through recognition patterns of HLA-DRB1*03:1, HLA- DRB1*07:1 and HLA-DRB1*15:1 (Dasgupta, unpublished observations) through computer guided molecular docking process. We extended our findings to address the question: What is the specificity of localized mucosal layer immune responses to the SARS-COV2 and its variants' Spike epitopes in human population? We seek explanation which will lead us to find out specific entry mechanisms of SARS-COV2 and its variants via first layer mucosal epithelial cells of respiratory tract, gastrointestinal tract and urogenital tract of human beings. Susceptibility is also a question as, within a population, not everyone acquired infection with same degree of severity to moribund or mortality. In this aspect, the degree of HLA-DRB1 alleles recognition to virus epitopes and MAIT-TCR response in a population have immense role towards localized inflammation leading severity.
Our findings showed specific interactions with binding affinity of mucosal associated invariant T cell-receptors (MAIT-TCR) for HLA-DRB1 allele -screened N-terminal Spike epitopes. The amino acid sequence specificity of the epitopes with MAIT- TCR is determined by molecular docking and molecular dynamics experiments.
Mucosal-associated invariant T cells are separate pool of lymphocytes belong to cytotoxic (CD8) and natural killer T cell (NKT) population in human which are known to recognize human Major histocompatibility Complex Class I related protein 1 (MR 1) and CD1d (23), (24), (16). The innate like T cells reside within tissues and share characteristics with alpha-beta and gamma- delta T cells. The antiviral effect of MAIT cells has been found in hepatitis B virus related hepatocellular carcinoma (25). Recent investigations demonstrated active role of MAIT cells in anti-tumor cell therapy approach for hematological malignancy (26), inflammation in central nervous system (CNS) during multiple sclerosis (MS) (27). Wang et al. (28) demonstrated presence of TRAV1- $^{2+}$ CD8 $^{+}$ MAIT cells and NCAM $^{hi}$ CD160 $^{+}$ NK cells in asymptomatic COVID-19 patients. These findings relate presence of innate-like MAIT cells during inflammatory responses in different tissues. The epitope specificity of SARS-COV2 and variants are found for innate immune responder TLR8 during versatile immune responses including brain (29). The observations presented here demonstrated critical role of HLA-DRB1 $^{*}$ 03:1, 07:1 and 15:1 recognition pattern for N-terminal Spike epitopes of SARS-COV2 and its variants Delta and Omicron for optimum immunogenicity which further led to sequence specific response by MAIT-TCR. Overall, experiments demonstrate recognition of the selected viral epitopes by population-predominant HLA-DRB1 alleles is a prime immunological phenomenon which in turn activates localized invariant T (MAIT) cells via MAIT-TCR to induce antiviral defense during infection.
### ACKNOWLEDGEMENT
I (SDG) acknowledge RNIRISC and Neuro Drug Research LLC core organization fund, Discover private fund to NeuroDrug Research.
1. Amirfakhryan H, Safari F. Outbreak of SARS-CoV2: Pathogenesis of infection and cardiovascular involvement. Hellenic J Cardiol.
2020.
2. Belot A, Antona D, Renolleau S, Javouhey E, Hentgen V, Angoulvant F, et al. SARS-CoV-2-related paediatric inflammatory multisystem syndrome, an epidemiological study, France, 1 March to 17 May 2020. Euro Surveill. 2020; 25(22).
3. Pan F, Yang L, Li Y, Liang B, Li L, Ye T, et al. Factors associated with death outcome in patients with severe coronavirus disease-19 (COVID-19): a case-control study. Int J Med Sci. 2020; 17(9): 1281-92.
4. De Luca G, Nardin M, Algowhary M, Uguz B, Oliveira DC, Ganyukov V, et al. Impact of chronic obstructive pulmonary disease on short-term outcome in patients with ST-elevation myocardial infarction during COVID-19 pandemic: insights from the international multicenter ISACS-STEMI registry. Respir Res. 2022; 23(1): 207.
5. Bhattacharya M, Chatterjee S, Sharma AR, Lee SS, Chakraborty C. Delta variant (B.1.617.2) of SARS-CoV-2: current understanding of infection, transmission, immune escape, and mutational landscape. Folia Microbiol (Praha).
2022.
6. Ribeiro Xavier C, Sachetto Oliveira R, da Fonseca Vieira V, Lobosco M, Weber Dos Santos R. Characterisation of Omicron Variant during COVID-19 Pandemic and the Impact of Vaccination, Transmission Rate, Mortality, and Reinfection in South Africa, Germany, and Brazil. BioTech (Basel). 2022; 11(2).
7. Xue L, Jing S, Zhang K, Milne R, Wang H. Infectivity versus fatality of SARS-CoV-2 mutations and influenza. Int J Infect Dis. 2022; 121: 195-202.
8. Antonio EC, Meireles MR, Bragatte MAS, Vieira GF. Viral immunogenic footprints conferring T cell cross-protection to SARS-CoV-2 and its variants. Front Immunol. 2022; 13: 931372.
9. Szeto C, Nguyen AT, Lobos CA, Chatzileontiadou DSM, Jayasinghe D, Grant EJ, et al. Molecular Basis of a Dominant SARS-CoV-2 Spike-Derived Epitope Presented by HLA- $A^{*}02:01$ Recognised by a Public TCR. Cells. 2021; 10(10).
10. Wragg KM, Lee WS, Koutsakos M, Tan HX, Amarasena T, Reynaldi A, et al. Establishment and recall of SARS-CoV-2 spike epitope-specific CD4(+) T cell memory. Nat Immunol. 2022; 23(5): 768-80.
11. Mairpady Shambat S, Gomez-Mejia A, Schweizer TA, Huemer M, Chang CC, Acevedo C, et al. Hyperinflammatory environment drives dysfunctional myeloid cell effector response to bacterial challenge in COVID-19. PLoS Pathog. 2022; 18(1): e1010176.
12. Payen D, Cravat M, Maadadi H, Didelot C, Prosic L, Dupuis C, et al. A Longitudinal Study of Immune
- Cells in Severe COVID-19 Patients. Front Immunol. 2020; 11: 580250.
13. Bonnet B, Cosme J, Dupuis C, Coupez E, Adda M, Calvet L, et al. Severe COVID-19 is characterized by the co-occurrence of moderate cytokine inflammation and severe monocyte dysregulation. EBioMedicine. 2021; 73: 103622.
14. Dobrijevic Z, Gligorijevic N, Sunderic M, Penezic A, Miljus G, Tomic S, et al. The association of human leucocyte antigen (HLA) alleles with COVID-19 severity: A systematic review and meta-analysis. Rev Med Virol. 2022;e2378.
15. Dhanda SK, Mahajan S, Paul S, Yan Z, Kim H, Jespersen MC, et al. IEDB-AR: immune epitope database-analysis resource in 2019. Nucleic Acids Res. 2019; 47(W1): W502-W6.
16. Harly C, Robert J, Legoux F, Lantz O. gammadelta T, NKT, and MAIT Cells During Evolution: Redundancy or Specialized Functions? J Immunol. 2022; 209(2): 217-25.
17. Johnson DN, Ruan Z, Petley EV, Devi S, Holz LE, Uldrich AP, et al. Differential location of NKT and MAIT cells within lymphoid tissue. Sci Rep. 2022; 12(1): 4034.
18. Dias J, Boulouis C, Gorin JB, van den Biggelaar R, Lal KG, Gibbs A, et al. The CD4(-)CD8(-) MAIT cell subpopulation is a functionally distinct subset developmentally related to the main CD8(+) MAIT cell pool. Proc Natl Acad Sci USA. 2018; 115(49): E11513-E22.
19. Kurioka A, Jahun AS, Hannaway RF, Walker LJ, Fergusson JR, Sverremark-Ekstrom E, et al. Shared and Distinct Phenotypes and Functions of Human CD161++ Valpha7.2+ T Cell Subsets. Front Immunol. 2017; 8: 1031.
20. Chiba A, Murayama G, Miyake S. Characteristics of mucosal-associated invariant T cells and their roles in immune diseases. Int Immunol. 2021; 33(12): 775-80.
21. Henin D, Pellegrini G, Carmagnola D, Lanza Attisano GC, Lopez G, Ferrero S, et al. Morphological and Immunopathological Aspects of Lingual Tissues in COVID-19. Cells. 2022; 11(7).
22. Slomka A, Martucci G, Raffa GM, Malvindi PG, Zekanowska E, Lorusso R, et al. Immunological and Hematological Response in COVID-19. Adv Exp Med Biol. 2021; 1352: 73-86.
23. Souter MNT, Awad W, Li S, Pedionco TJ, Meehan BS, Meehan LJ, et al. CD8 coreceptor engagement of MR1 enhances antigen responsiveness by human MAIT and other MR1-reactive T cells. J Exp Med. 2022; 219(9).
24. Awad W, Le Nours J, Kjer-Nielsen L, McCluskey J, Rossjohn J. Mucosal-associated invariant T cell receptor recognition of small molecules presented by MR1. Immunol Cell Biol. 2018; 96(6): 588-97.
25. Healy K, Pavesi A, Parrot T, Sobkowiak MJ, Reinsbach SE, Davanian H, et al. Human MAIT cells endowed with HBV specificity are cytotoxic and migrate towards HBV-HCC while retaining antimicrobial functions. JHEP Rep. 2021; 3(4): 100318.
26. Gao MG, Zhao XS. Mining the multifunction of mucosal-associated invariant T cells in hematological malignancies and transplantation immunity: A promising hexagon soldier in immunomodulatory. Front Immunol. 2022; 13: 931764.
27. Gargano F, Guerrera G, Piras E, Serafini B, Di Paola M, Rizzetto L, et al. Proinflammatory mucosal
- associated invariant CD8+ T cells react to gut flora yeasts and infiltrate multiple sclerosis brain. Front Immunol. 2022; 13: 890298.
28. Wang X, Bai H, Ma J, Qin H, Zeng Q, Hu F, et al. Identification of Distinct Immune Cell Subsets Associated With Asymptomatic Infection, Disease Severity, and Viral Persistence in COVID-19 Patients. Front Immunol. 2022; 13: 812514.
29. Dasgupta S, Bandyopadhyay M. Molecular docking of SARS-COV-2 Spike epitope sequences identifies heterodimeric peptide-protein complex formation with human Zo-1, TLR8 and brain specific glial proteins. Med Hypotheses. 2021; 157: 110706.
Table 1: Epitope peptide sequence- protein database files
<table><tr><td>Protein Database (PDB) files</td><td>Epitope inserted peptide sequences</td></tr><tr><td>OM42Omicron Spike (B.1.1.529)</td><td>SSQCVNLTTRTQLPPAYTNSFTRGVYYPDK(SEQ. 12- 30)</td></tr><tr><td>OM60Omicron Spike (B.1.1.529)</td><td>VFRSSVLHSTQDLFLPFFSNVTWFHVISGT(SEQ. 31-60)</td></tr><tr><td>M312Omicron Spike (B.1.1.529)</td><td>TSNFRVQPTESIVRFPNITNLCPFDEVFNAT(SEQ. 312-342)</td></tr><tr><td>M370Omicron Spike (B.1.1.529)</td><td>RFASVYAWNRKRISNCVADYSVLYNAPF(SEQ. 343- 371)</td></tr><tr><td>DLT1Delta Spike (B.1.617.2)</td><td>FASVYAWNR(SEQ.24- 32)</td></tr><tr><td>DLT2SARS-COV2 and Delta Spike(B.1.617.2)</td><td>LYNSASFSTDelta Spike (7ORB_R: SEQ.45-53)SARS-COV2 (6VXX_A: SEQ: 397-405)</td></tr></table>
Table 2: HLA-DRB1 recognition of epitope peptides from first thousand N- terminal amino acid sequences of SARS-COV2 Omicron variant B. 1.1.529 (7QO9_A).
<table><tr><td rowspan="2">Epitope Peptides</td><td rowspan="2">Length</td><td colspan="3">HLA-DRB1*</td></tr><tr><td>03:01</td><td>07:01</td><td>15:01</td></tr><tr><td>MFVFLVLLPLVSSQCVN</td><td>1-17</td><td>74.94</td><td>52.80</td><td>5.11</td></tr><tr><td>MFVFLVLLPLVSSQCVNL</td><td>1-18</td><td>76.95</td><td>73.99</td><td>9.77</td></tr><tr><td>FVFLVLLPLVSSQCVNL</td><td>2-18</td><td>40.88</td><td>37.47</td><td>5.11</td></tr><tr><td>VLLPLVSSQCVNLTTRQ</td><td>6-23</td><td>79.91</td><td>106.55</td><td>59.19</td></tr><tr><td>LLPLVSSQCVNLTTR</td><td>7-21</td><td>17</td><td>28</td><td>21</td></tr><tr><td>LLPLVSSQCVNLTTRTQL</td><td>7-24</td><td>65.11</td><td>79.91</td><td>62.15</td></tr><tr><td>LPLVSSQCVNLTTRTQLP</td><td>8-25</td><td>65.11</td><td>121.35</td><td>68.07</td></tr><tr><td>PLVSSQCVNLTTRTQL</td><td>9-24</td><td>20.78</td><td>39.25</td><td>61.18</td></tr></table>
SARS-COV2 Omicron variant peptides (selected from the first thousand N terminal amino acid sequence) are analyzed by Immune database Tools (IEDB Tools) for their HLA-DRB1 allele recognition and CD4 immunogenic recognition ability. The lowest score indicates the most effective towards immunogenic response.
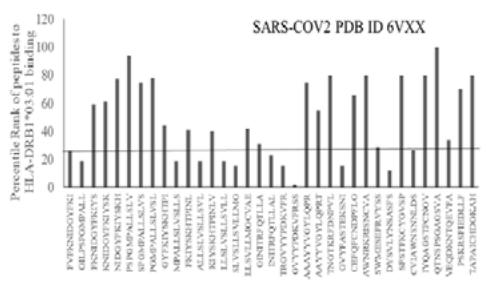

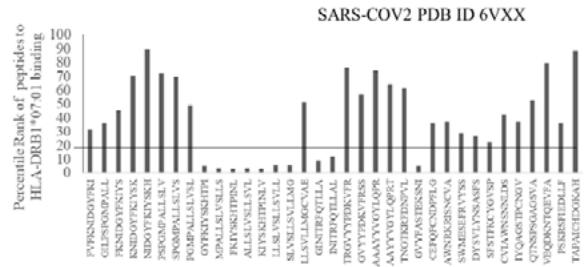
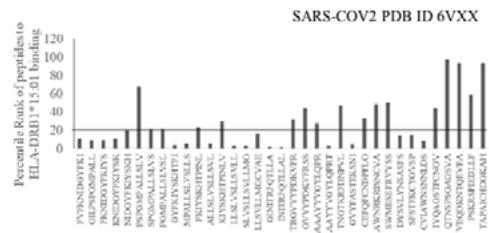
Fig.1A
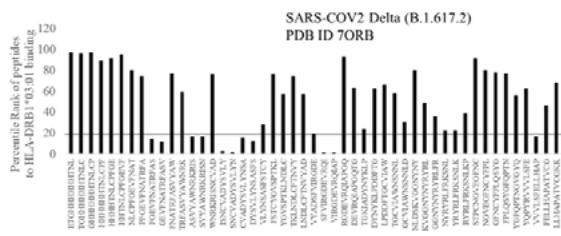



Fig.1B Fig.1 A, B: Selection of immunogenic T cell specific epitopes by HLA-DRB1 alleles from Spike envelop protein of (A) SARS-COV2 (6VXX_A) and (B) Delta variant (B.1.617.2) virus. The Spike envelop proteins are selected from protein database. The 60 amino acid peptide strings are processed by Immune Epitope Database (IEDB) Tool software system. The tool was programmed for detecting 12 amino acid epitope peptides responding to HLA-DRB1*03:01, HLA-DRB1*07:01 and HLA-DRB1*15:01 alleles representing population based immunogenic responses. (A) Immunogenic Spike epitopes from SARS-COV2 6VXX_A protein are mapped on the basis of percentile rank determining binding affinity to HLA-DRB1. The base line of 20 percentile range is shown in the plot. The epitopes below this line are low immunogenic and good fit to corresponding HLA-DRB1 alleles. (B) Immunogenic Spike epitopes from Delta variant (B.1.617.2) are mapped in a similar way and scaled through 20 percentile base line. The low rank immunogenic peptide epitopes are found below 20 percentile rank line. Each bar in the plot represents percentile HLA-DRB1 alleles 03:01, 07:01, 15:01 responses to each of the peptide epitopes presented in the figure.
1 6 11 16 21 26 31 36 41 46 51 56 61 66 71 76 81 86 91 96 101 106 111 116 121 126 IQRPPKIOV SRHPPEDGKPNYLCNYYGFHPQIEIDLLKNGEKIKSEQDSLSFSKDMSFYLLSRAFTPNKSDDYSCRVKHTLEQPRIVKMDRDLGGGGSGGSGGGSRTHSLRYFRLGISEPGY LYNSSFST

Delta epitope DLT2 versus MAIT TCR
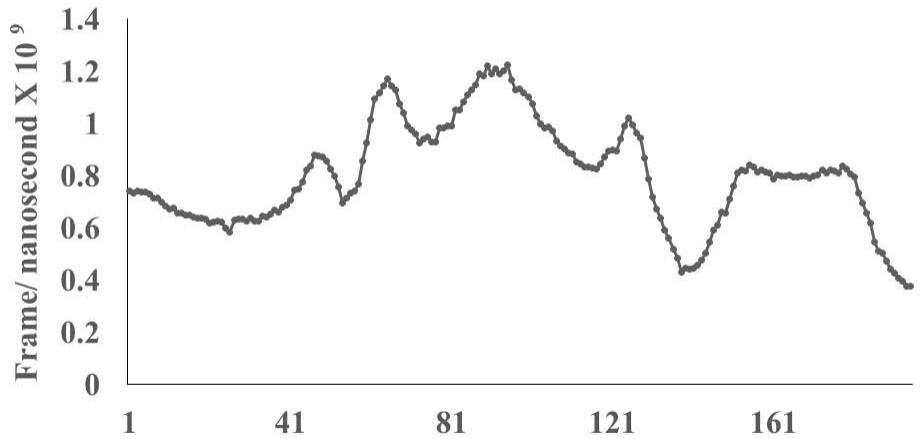
RMSD: 2.309 Angstrom Fig.2B Rendering Time X $10^{9}$ nanosecond
/4L9L/A/C 16 11 16 21 26 31 36 41 46 51 56 61 66 71 76 81 86 91 96 101 106 111 116 121 IQRPPKIQQVY SRHPPEDGKPNYLNCVYVGFHPQIEIDLLKNGEIKSEQDSLSFKDMSFYLLSHEFTPNSKDQYSCRVKHTLEQPRVKMDRLGGGGSGSGGGSRTHSLRYFRLGI 16 FASVYAUNR
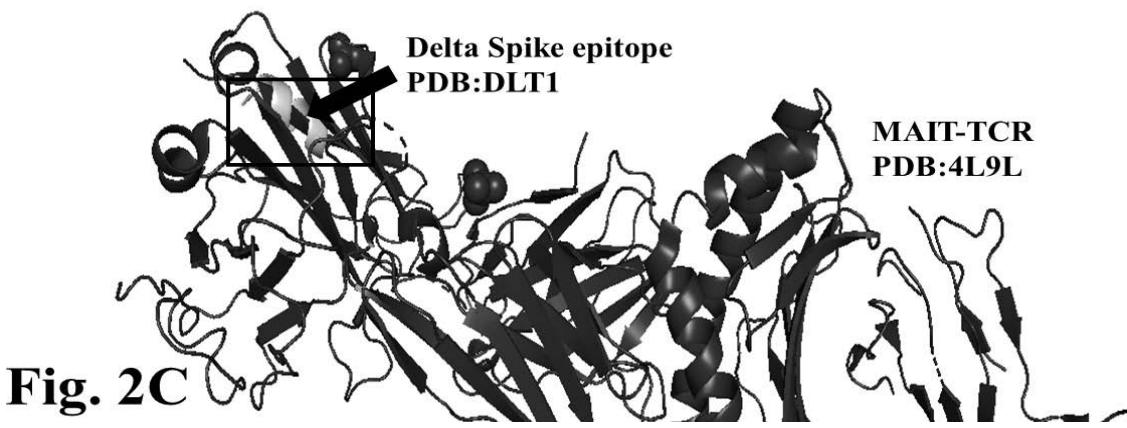
DLT1 versus MAIT-TCR
RMSD: 5.410 Angstrom
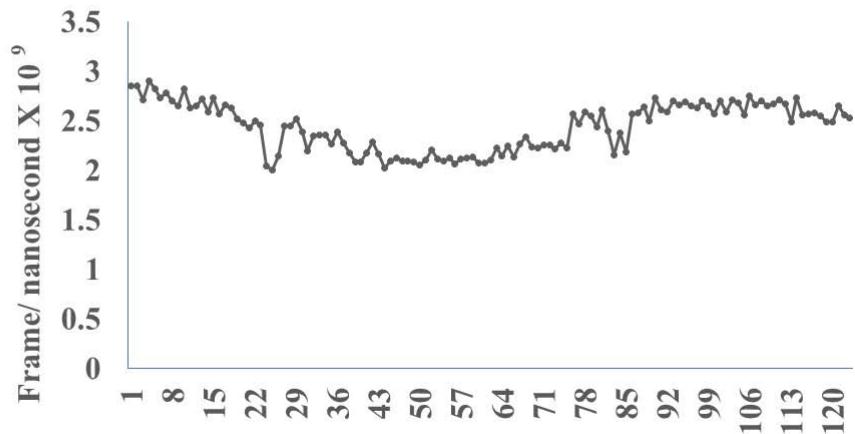
Fig 2 (A-D) Statistical significance $(\mathrm{p} < 0.05)$ for binding and close atomic interactions was determined by one factor ANOVA between Frame per nanosecond (Angstrom) value of identified SARSCOV2 Delta N terminal spike epitopes Spike epitopes and Control unrelated Mycobacterium avium antigen 85 peptide. Results obtained from Molecular Dynamics experiments (Fig 2A to D).
Fig. 2D
Rendering Time (nanosecond) X $10^{9}$
Fig.2 A, B, C, D: Determination of specific binding of immunogenic Spike epitope sequences from SARS-COV2 and Delta variant (B.1.617.2) with mucosa-associated invariant T cell receptor (MAIT-TCR). The specific binding patterns between the closest aligned atoms within cut off distance 2 Angstrom are determined by 10 cycle Molecular Dynamics alignment assay using epitopes as mobile unit and MAIT-TCR is the target for none to one interaction. The root mean square deviation (RMSD, Angstrom) for each experiment between epitope peptide and MAIT-TCR demonstrate good fit alignment. The lesser RMSD, the better is the alignment thus represents better binding. The protein database files (PDB) are used in the experiments. (A) The molecular docking of Delta variant epitope DLT2 (shown by arrow) binds with MAIT-TCR. The amino acid sequence specific alignment is shown in a box at the top of the figure. The binding area is also shown by a box in the figure. (B) The molecular dynamics experiment demonstrates binding pattern of interacting atoms between DLT2 (PDB) and MAIT-TCR (PDB: 4L9L). RMSD: 2.309 Angstrom. (C) The molecular docking of Delta Spike epitope (DLT1) with MAIT-TCR (PDB: 4L9L). The interacting protein sequences are shown in a box at the top of the figure. The binding of DLT1 by MAIT-TCR is shown in a box in the figure. (D) The molecular dynamics experiment demonstrates binding pattern of interacting atoms between DLT1 (PDB) and MAIT-TCR (PDB: 4L9L). RMSD: 5.410 Angstrom. All these interaction events in the Molecular Dynamics experiments are based on oscillation at the rate of 30 frame per nanosecond for 100 nanosecond duration as described in Materials and Methods. Statistical significance $(p < 0.05)$ of the frame per nanosecond values of experimental epitope peptides as compared with unrelated control peptide antigen85 with respect to MAIT-TCR has been determined by one factor ANOVA.
Anova: Single Factor
<table><tr><td colspan="5">SUMMARY</td></tr><tr><td>Groups</td><td>Count</td><td>Sum</td><td>Average</td><td>Variance</td></tr><tr><td>Control</td><td>42</td><td>35.47787</td><td>0.844711</td><td>0.00052</td></tr><tr><td>DLT1</td><td>42</td><td>104.2915</td><td>2.483131</td><td>0.061207</td></tr><tr><td>DLT2</td><td>42</td><td>27.92395</td><td>0.664856</td><td>0.001887</td></tr></table>
<table><tr><td colspan="7">ANOVA</td></tr><tr><td>Source of Variation</td><td>SS</td><td>df</td><td>MS</td><td>F</td><td>P-value</td><td>F crit</td></tr><tr><td>Between Groups</td><td>84.32049</td><td>2</td><td>42.16024</td><td>1988.244</td><td>2.22E-94</td><td>3.069894</td></tr><tr><td>Within Groups</td><td>2.608186</td><td>123</td><td>0.021205</td><td></td><td></td><td></td></tr><tr><td>Total</td><td>86.92867</td><td>125</td><td></td><td></td><td></td><td></td></tr></table>
<table><tr><td>/4L9L/A/C</td><td>1</td><td>6</td><td>11</td><td>16</td><td>21</td><td>26</td><td>31</td><td>36</td><td>41</td><td>46</td><td>51</td><td>56</td><td>61</td><td>66</td><td>71</td><td>76</td><td>81</td><td>86</td><td>91</td><td>96</td><td>101</td><td>106</td><td>111</td><td>116</td><td>121</td><td>126</td><td></td></tr><tr><td colspan="2">IQRPPKIQVYSRHPEDGKPNLYNCVYYGF</td><td>1</td><td>6</td><td>11</td><td>16</td><td>21</td><td>26</td><td>31</td><td>36</td><td>41</td><td>46</td><td>51</td><td>56</td><td>61</td><td>66</td><td>71</td><td>76</td><td>81</td><td>86</td><td>91</td><td>96</td><td>101</td><td>106</td><td>111</td><td>116</td><td>121</td><td>126</td></tr><tr><td>/OM42//A</td><td colspan="2">SSOCVNLTTRTQLPPRTNSFTRGVYYPDK</td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td></tr></table>
<table><tr><td>/4L9L/A/C</td><td>1</td><td>6</td><td>11</td><td>16</td><td>21</td><td>26</td><td>31</td><td>36</td><td>41</td><td>46</td><td>51</td><td>56</td><td>61</td><td>66</td><td>71</td><td>76</td><td>81</td><td>86</td><td>91</td><td>96</td><td>101</td><td>106</td><td>111</td><td>116</td><td>121</td><td>126</td><td></td></tr><tr><td colspan="2">IQRPPKIQVYSRHPEDGKPNLYNCVYYGF</td><td>1</td><td>6</td><td>11</td><td>16</td><td>21</td><td>26</td><td>31</td><td>36</td><td>41</td><td>46</td><td>51</td><td>56</td><td>61</td><td>66</td><td>71</td><td>76</td><td>81</td><td>86</td><td>91</td><td>96</td><td>101</td><td>106</td><td>111</td><td>116</td><td>121</td><td>126</td></tr><tr><td>/OM42//A</td><td colspan="2">SSOCVNLTTRTQLPPRTNSFTRGVYYPDK</td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td></tr></table>
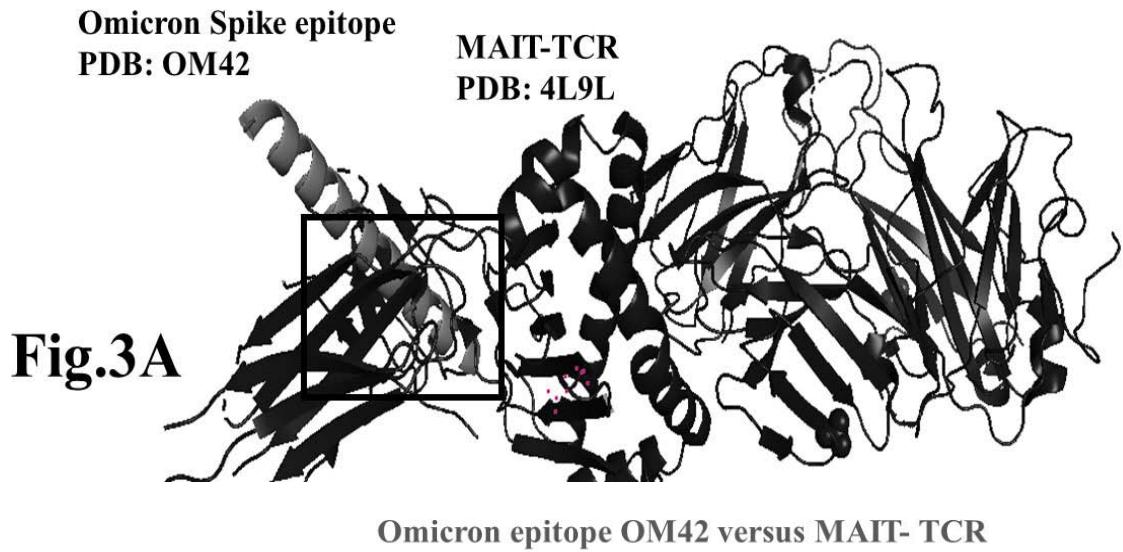
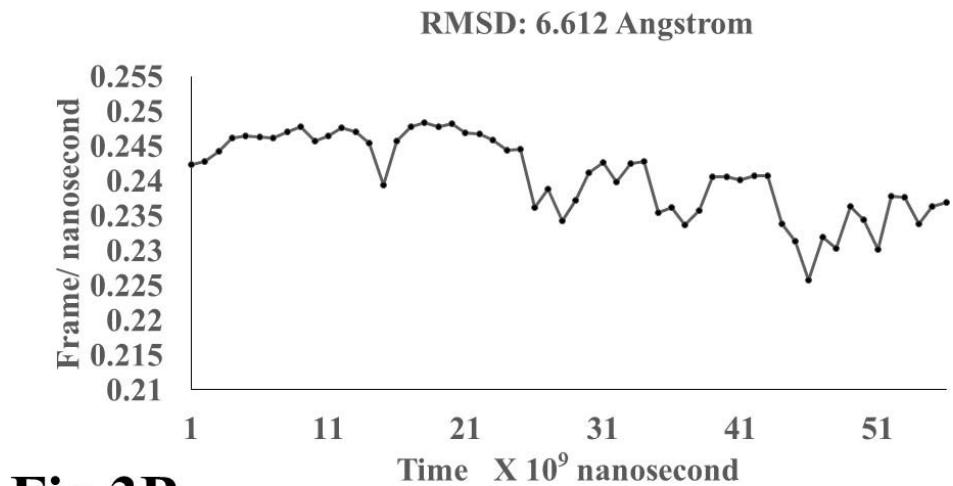
Fig.3B
Omicron epitope OM60 vs. MAIT-TCR
RMSD: 7.282 Angstrom Fig.3D
Omicron epitope M312 vs MAIT-TCR
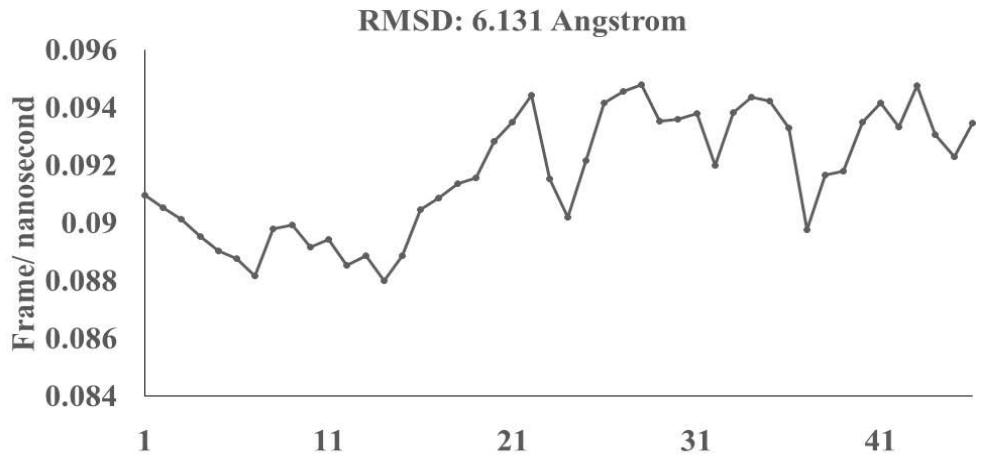
Fig.3F Rendering Time X 10 $^{9}$ nanosecond
<table><tr><td rowspan="2">/4L9L/R/D</td><td>1</td><td>6</td><td>11</td><td>16</td><td>21</td><td>26</td><td>31</td><td>36</td><td>41</td><td>46</td><td>51</td><td>56</td><td>61</td><td>66</td><td>71</td><td>76</td><td>81</td><td>86</td><td>91</td><td>96</td><td>101</td><td>106</td><td>111</td><td>116</td><td>121</td><td>126</td><td>131</td><td></td></tr><tr><td colspan="6">IQRPPKIQVYSRHPPEDGKPNYLCNY/N</td><td colspan="22">GFHPQQIEIDLKNGEIKSEQSDLSFSKDMSFYLLSHAETPNSKIDQYSCRVKHTLEQPRIVKMJDRDLGGGGSGGGSGGGGSRTHSLRYFRLGISEPGYGIP</td></tr><tr><td rowspan="2">/M370//A</td><td>1</td><td>6</td><td>11</td><td>16</td><td>21</td><td>26</td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td></tr><tr><td colspan="6">RFASVYAMNRKRISNCVADYSVLYNLAPF</td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td><td></td></tr></table>
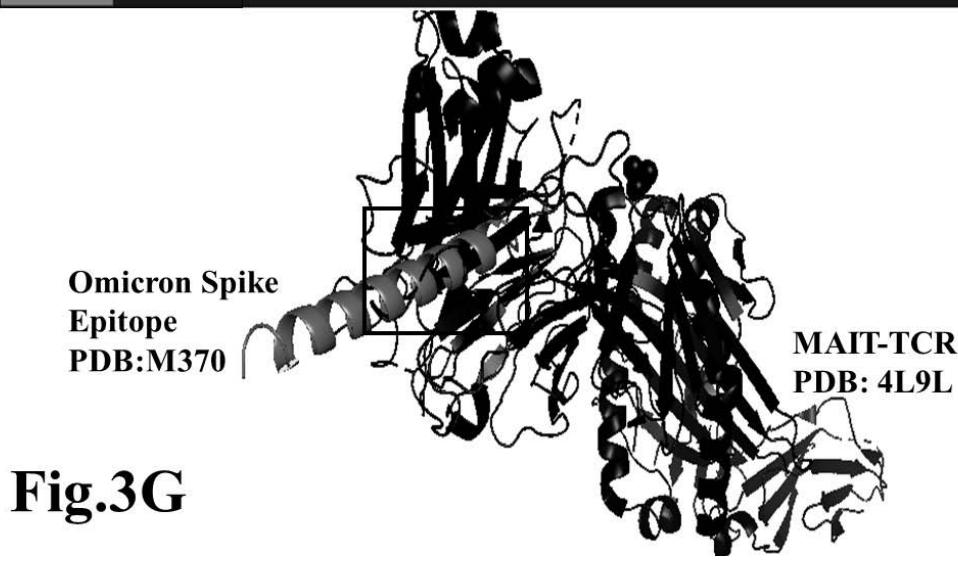
Fig.3G
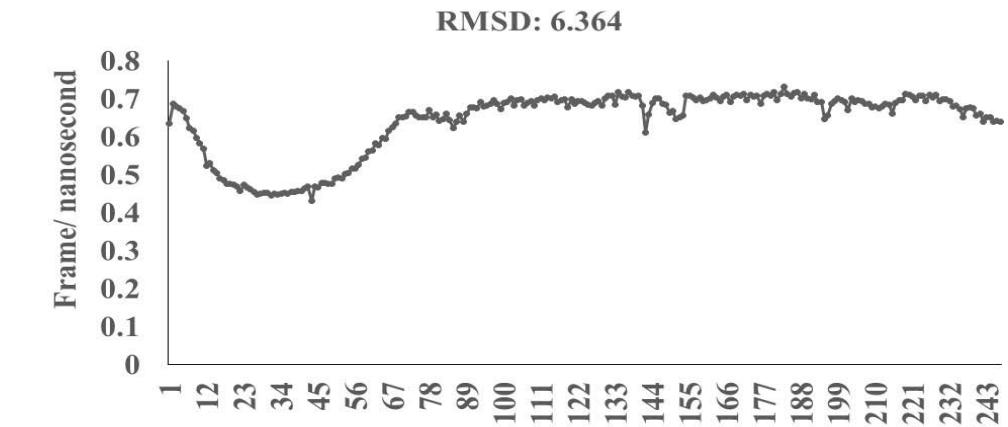
Omicron epitope M370 vs MAIT TCR Fig.3H Rendering Time X $10^{9}$ nanosecond Fig.3 A, B, C, D, E, F, G, H: Epitope peptides of Omicron variant (B.1.1.529) bind with mucosa-associated invariant T cell receptor (MAIT-TCR). The molecular docking and molecular dynamics experiments are performed to determine specific binding nature of Omicron epitopes and MAIT-TCR. The closest aligned atoms of amino acids within the cut off distance 2 Angstrom are determined by 10 cycle Molecular Dynamics alignment assay using the epitopes as mobile unit and MAIT-TCR is the target for none to one interaction. The root mean square deviation (RMSD, Angstrom) for each experiment between epitope peptide and MAIT-TCR demonstrate good fit alignment. The lesser RMSD, the better is the alignment thus represents better binding. The protein database files (PDB) are used in the experiments. (A) The molecular docking of Omicron variant Spike epitope (PDB: OM42) with MAIT-TCR (PDB: 4L9L). The interacting protein sequences are shown in a box at the top of the figure. The binding of OM42 by MAIT-TCR is shown in a box in the figure. (B) The molecular dynamics experiment demonstrates binding pattern of interacting atoms between OM42 (PDB) and MAIT-TCR (PDB: 4L9L). RMSD: 6.612 Angstrom. (C) The molecular docking of Omicron variant Spike epitope (PDB: OM60) with MAIT-TCR (PDB: 4L9L). The interacting protein sequences are shown in a box at the top of the figure. The binding of OM60 by MAIT-TCR is shown in a box in the figure. (D) The molecular dynamics experiment demonstrates binding pattern of interacting atoms between OM60 (PDB) and MAIT-TCR (PDB: 4L9L). RMSD: 7.282 Angstrom. The RMSD value shows comparatively less aligned amino acid sequences are present in OM60 epitope in the aspect of interaction with MAIT-TCR. (E) Molecular docking of Omicron variant Spike epitope (PDB: M312) with MAIT-TCR (PDB: 4L9L). The interacting protein sequences are shown in a box at the top of the figure. The binding zone of epitope peptide with MAIT-TCR is shown in a box in the figure. (F) The molecular dynamics experiment demonstrates binding of interacting atoms between M312 (PDB) and MAIT-TCR (PDB: 4L9L). RMSD: 6.131 Angstrom. (G) Molecular docking of Omicron variant Spike epitope (PDB:M370) with MAIT-TCR (PDB: 4L9L). The interacting protein sequences are shown in a box at the top of the figure. The binding of M370 by MAIT-TCR is shown in a box in the figure. (H) Molecular dynamics experiment demonstrates binding pattern of interacting atoms between epitope M370 (PDB) and MAIT-TCR (PDB: 4L9L). RMSD: 6.364 Angstrom. All these interaction events in the Molecular Dynamics experiments are based on oscillation atoms of interacting proteins defined at the rate of 30 frame per nanosecond for 100 nanosecond duration as described in Materials and Methods. All results are expressed as Frame per nanosecond versus Rendering time (nanosecond) at a particular executive root mean square deviation (RMSD) value. Statistical significance $(p < 0.05)$ of the frame per nanosecond values of experimental epitope peptides as compared with unrelated control peptide antigen85 with respect to MAIT-TCR has been determined by one factor ANOVA.
SUMMARY
<table><tr><td>Groups</td><td>Count</td><td>Sum</td><td>Average</td><td>Variance</td></tr><tr><td>Control</td><td>42</td><td>35.47787</td><td>0.844711</td><td>0.00052</td></tr><tr><td>OM42</td><td>42</td><td>10.20515</td><td>0.24298</td><td>1.87E-05</td></tr><tr><td>OM60</td><td>42</td><td>25.81601</td><td>0.614667</td><td>0.006575</td></tr><tr><td>M312</td><td>42</td><td>3.844725</td><td>0.091541</td><td>4.45E-06</td></tr><tr><td>M370</td><td>42</td><td>21.48756</td><td>0.511609</td><td>0.006286</td></tr></table>
ANOVA
<table><tr><td>Source of Variation</td><td>SS</td><td>df</td><td>MS</td><td>F</td><td>P-value</td><td>F crit</td></tr><tr><td>Between Groups</td><td>15.01255</td><td>4</td><td>3.753137</td><td>1399.918</td><td>1.5E-147</td><td>2.415694</td></tr><tr><td>Within Groups</td><td>0.549599</td><td>205</td><td>0.002681</td><td></td><td></td><td></td></tr><tr><td>Total</td><td>15.56215</td><td>209</td><td></td><td></td><td></td><td></td></tr></table>
Generating HTML Viewer...
References
24 Cites in Article
H Amirfakhryan,F Safari (2020). Outbreak of SARS-CoV2: Pathogenesis of infection and cardiovascular involvement.
Alexandre Belot,Denise Antona,Sylvain Renolleau,Etienne Javouhey,Véronique Hentgen,François Angoulvant,Christophe Delacourt,Xavier Iriart,Caroline Ovaert,Brigitte Bader-Meunier,Isabelle Kone-Paut,Daniel Levy-Bruhl (2020). SARS-CoV-2-related paediatric inflammatory multisystem syndrome, an epidemiological study, France, 1 March to 17 May 2020.
Feng Pan,Lian Yang,Yuncheng Li,Bo Liang,Lin Li,Tianhe Ye,Lingli Li,Dehan Liu,Shan Gui,Yu Hu,Chuansheng Zheng (2020). Factors associated with death outcome in patients with severe coronavirus disease-19 (COVID-19): a case-control study.
G De Luca,M Nardin,M Algowhary,B Uguz,D Oliveira,V Ganyukov (2022). Impact of chronic obstructive pulmonary disease on short-term outcome in patients with ST-elevation myocardial infarction during COVID-19 pandemic: insights from the international multicenter ISACS-STEMI registry.
Manojit Bhattacharya,Srijan Chatterjee,Ashish Sharma,Sang-Soo Lee,Chiranjib Chakraborty (2022). Delta variant (B.1.617.2) of SARS-CoV-2: current understanding of infection, transmission, immune escape, and mutational landscape.
Ribeiro Xavier,C,Sachetto Oliveira,R Da,Fonseca Vieira,V Lobosco,M,Weber Santos,R (2022). Characterisation of Omicron Variant during COVID-19 Pandemic and the Impact of Vaccination, Transmission Rate, Mortality, and Reinfection in South Africa, Germany, and Brazil.
Ling Xue,Shuanglin Jing,Kai Zhang,Russell Milne,Hao Wang (2022). Infectivity versus fatality of SARS-CoV-2 mutations and influenza.
Eduardo Antonio,Mariana Meireles,Marcelo Bragatte,Gustavo Vieira (2022). Viral immunogenic footprints conferring T cell cross-protection to SARS-CoV-2 and its variants.
Christopher Szeto,Andrea Nguyen,Christian Lobos,Demetra Chatzileontiadou,Dhilshan Jayasinghe,Emma Grant,Alan Riboldi-Tunnicliffe,Corey Smith,Stephanie Gras (2021). Molecular Basis of a Dominant SARS-CoV-2 Spike-Derived Epitope Presented by HLA-A*02:01 Recognised by a Public TCR.
K Wragg,W Lee,M Koutsakos,H Tan,T Amarasena,A Reynaldi (2022). Establishment and recall of SARS-CoV-2 spike epitope-specific CD4(+) T cell memory.
Didier Payen,Maxime Cravat,Hadil Maadadi,Carole Didelot,Lydia Prosic,Claire Dupuis,Marie-Reine Losser,Marcelo De Carvalho Bittencourt (2020). A Longitudinal Study of Immune Cells in Severe COVID-19 Patients.
Benjamin Bonnet,Justine Cosme,Claire Dupuis,Elisabeth Coupez,Mireille Adda,Laure Calvet,Laurie Fabre,Pierre Saint-Sardos,Marine Bereiziat,Magali Vidal,Henri Laurichesse,Bertrand Souweine,Bertrand Evrard (2021). Severe COVID-19 is characterized by the co-occurrence of moderate cytokine inflammation and severe monocyte dysregulation.
Zorana Dobrijević,Nikola Gligorijević,Miloš Šunderić,Ana Penezić,Goran Miljuš,Sergej Tomić,Olgica Nedić (2022). The association of human leucocyte antigen (HLA) alleles with COVID‐19 severity: A systematic review and meta‐analysis.
S Dhanda,S Mahajan,S Paul,Z Yan,H Kim,M Jespersen (2019). IEDB-AR: immune epitope database-analysis resource in 2019.
Christelle Harly,Jacques Robert,Francois Legoux,Olivier Lantz (2022). γδ T, NKT, and MAIT Cells During Evolution: Redundancy or Specialized Functions?.
Darryl Johnson,Zheng Ruan,Emma Petley,Sapna Devi,Lauren Holz,Adam Uldrich,Jeffrey Mak,Jyh Hor,Scott Mueller,James Mccluskey,David Fairlie,Phillip Darcy,Paul Beavis,William Heath,Dale Godfrey (2022). Differential location of NKT and MAIT cells within lymphoid tissue.
Joana Dias,Caroline Boulouis,Jean-Baptiste Gorin,Robin Van Den Biggelaar,Kerri Lal,Anna Gibbs,Liyen Loh,Muhammad Gulam,Wan Sia,Sudipto Bari,William Hwang,Douglas Nixon,Son Nguyen,Michael Betts,Marcus Buggert,Michael Eller,Kristina Broliden,Annelie Tjernlund,Johan Sandberg,Edwin Leeansyah (2018). The CD4<sup>−</sup>CD8<sup>−</sup>MAIT cell subpopulation is a functionally distinct subset developmentally related to the main CD8<sup>+</sup>MAIT cell pool.
A Kurioka,A Jahun,R Hannaway,L Walker,J Fergusson,E Sverremark-Ekstrom (2017). Shared and Distinct Phenotypes and Functions of Human CD161++ Valpha7.2+ T Cell Subsets.
A Chiba,G Murayama,S Miyake (2021). Characteristics of mucosal-associated invariant T cells and their roles in immune diseases.
Dolaji Henin,Gaia Pellegrini,Daniela Carmagnola,Giuseppe Lanza Attisano,Gianluca Lopez,Stefano Ferrero,Antonella Amendola,Danilo De Angelis,Elisabetta Tanzi,Claudia Dellavia (2022). Morphological and Immunopathological Aspects of Lingual Tissues in COVID-19.
Artur Słomka,Gennaro Martucci,Giuseppe Raffa,Pietro Malvindi,Ewa Żekanowska,Roberto Lorusso,Piotr Suwalski,Mariusz Kowalewski (2021). Immunological and Hematological Response in COVID-19.
Michael Souter,Wael Awad,Shihan Li,Troi Pediongco,Bronwyn Meehan,Lucy Meehan,Zehua Tian,Zhe Zhao,Huimeng Wang,Adam Nelson,Jérôme Le Nours,Yogesh Khandokar,T Praveena,Jacinta Wubben,Jie Lin,Lucy Sullivan,George Lovrecz,Jeffrey Mak,Ligong Liu,Lyudmila Kostenko,Katherine Kedzierska,Alexandra Corbett,David Fairlie,Andrew Brooks,Nicholas Gherardin,Adam Uldrich,Zhenjun Chen,Jamie Rossjohn,Dale Godfrey,James Mccluskey,Daniel Pellicci,Sidonia Eckle (2022). CD8 coreceptor engagement of MR1 enhances antigen responsiveness by human MAIT and other MR1-reactive T cells.
Wael Awad,Jérôme Le Nours,Lars Kjer‐nielsen,James Mccluskey,Jamie Rossjohn (2018). Mucosal‐associated invariant T cell receptor recognition of small molecules presented by <scp>MR</scp>1.
Explore published articles in an immersive Augmented Reality environment. Our platform converts research papers into interactive 3D books, allowing readers to view and interact with content using AR and VR compatible devices.
Your published article is automatically converted into a realistic 3D book. Flip through pages and read research papers in a more engaging and interactive format.
Our website is actively being updated, and changes may occur frequently. Please clear your browser cache if needed. For feedback or error reporting, please email [email protected]
Thank you for connecting with us. We will respond to you shortly.