Interestingly, in sustainable crop protection, disease diagnosis, and management are crucial in sustainable crop production. It plays a captious role in rain-fed pulses because the occurrence of season of cropping, cultivation after main crop, availability of soil moisture in poor conditions, consecutively following the same cultivars are acting a predominant role in disease diagnosis approaches and confirmation. Under these situations, occurrence of the manual errors (or) mis find faults resulting in complete drawbacks to disease diagnosis and management for farmers and scientists worldwide. Keeping this background, applying deep learning techniques is most helpful in diagnosing plant diseases silently and superiorly. Deep learning techniques were carried out in this study to diagnose foliar diseases in black gram such as anthracnose, leaf crinkle, powdery mildew, and yellow mosaic that causes a severe yield loss (>50%) silently accompanied by green biomass.
## I. INTRODUCTION
Timely diagnosis of plant diseases plays an important role in crop protection and sustainable agriculture [1]. The past decade has seen advances in science in all fields, particularly in agriculture, including the detection of plant diseases, the development of resistant varieties, and the introduction of advanced management approaches [2]. Generally, the diagnosis of plant diseases is classified according to field and laboratory conditions, based on prevalence, symptoms and signs (In vivo), cultivation, and characterization (In vitro) [3]. Minor errors and misidentifications due to complex diseases can sometimes occur during early diagnosis under field conditions due to visual inspection, sampling, and manual errors by untrained people [4]. Most cases occurred with foliar diseases and non-cultivated pathogens, leading to errors in diagnosis and subsequent treatment. Under this situation, deep learning technology in agriculture can be used in various ways for its development [5]. These techniques include various types such as Convolutional Neural Networks (CNNs), Augmented Short-Term Memory Networks (LSTMs), Recurrent Neural Networks (R.N.N.), Generative Adversarial Networks (GANs), Radial Basis Function Networks (RBFNs), and Multilayer Perceptron's. the algorithm is used. (M.L.P.), Self-Organizing Maps (S.O.M.), Deep Belief Networks (DBN), Restricted Boltzmann Machines (R.B.M.), and Autoencoders [6]. These convolutional neural networks (CNNs) are used for irrigation, fertilization planning, monitoring plant growth trends, drought signs, pest infestation, plant disease detection and diagnosis, and crop assessment related to ripening and harvesting for sustainable agricultural production [7,8]. CNN (Convolutional Neural Network) is a unique and advanced class of Artificial Neural Networks (ANN), commonly used for visual image analysis, and primarily for extracting features from gridded matrix data sets. This includes input layers (containing raw images), convolutional layers (extracting features from the input dataset through learnable filters or kernels), and pooling layers (containing regularly inserted images such as transforms (neuron code), reduction in calculation amount of volume to prevent overfitting through typical layers of max and average pooling), a fully connected layer (computing the final classification and regression), and an output layer (taking the dataset from the previous layer), providing probability evaluation through sigmoid or softmax tasks) [9]. With the exception of image insertion, these functions are entirely automated.
Based on these auto-perception and rapid comparisons, computational technologies are useful in the early identification of plant diseases for improved management. [10, 11].
Nowadays, advanced deep learning techniques (D.L.T.s) have been used better in CNNs by implementing different types of architectures or models. Especially, AlexNet, GoogLeNet, ResNet (tomato leaf curl diseases), LeNet (Banana foliar diseases), AlexNetOWTbn, Overfeat, V.G.G., Xception, SqueezeNet, and VGG-Inception models. Among them, VGG-Inception outclassed all the other models. The comparison and superior class were scored between the models by performance metrics of optimization, customization, sensitivity, specificity, and F1-score [12]. This strategy was used to diagnose foliar diseases and disorders of crops in agriculture. In particular, high green-biomass-producing legume crops such as green gram, black gram, cowpea, lentil, pigeon pea, and peas were studied by deep learning techniques for better yield and management of diseases [13, 14]. Because their potential ability to yield depends on their vegetative growth, when it is affected, the yield is also reduced [15]. Among these legumes, black gram (Vigna mungo L.) is one of the most essential legume crops predominantly cultivated on rainfed conditions in India with an area of 44.9 lakh ha. and accounting for a production of 26.2 lakh tonnes. In Tamil Nadu, 3.72 lakh hectares with a production of 1,262 lakh metric tonnes are associated with a productivity of $645\mathrm{kg / ha}$ [16]. In particular, its yield has reduced due to fungal and viral diseases like powdery mildew, anthracnose, and yellow mosaic from 9.0 to $50.0\%$ during the vegetative to reproductive stage. The severity of these diseases also depends upon the stage of infection, virulence, genotypes, and environmental factors [17]. Furthermore, by approaching manual diagnosis, these factors cause faults or misleads in diagnosis under field conditions. It encourages the application of deep learning algorithms to diagnose and manage plant diseases [18]. Considering this background, the study was carried out to diagnose foliar diseases in black gram (Vigna mungo L.) using deep learning algorithms.
## II. MATERIALS AND METHODS
In this study, the work was carried out initially with a survey and collection of samples, application of advanced V3 inception model progresses such as preprocessing of images, learning phase and evaluation phase for diagnosis of foliar diseases in black gram.
### a) Survey and Collection of Disease Images
Depending upon the disease prevalence, consecutively cultivating of same varieties of rice fallow black gram, this location was selected for these studies. During Kharif 2022, a vast survey was conducted on rice fallow black gram cultivated fields in four different blocks, such as Aranthangi, Gandarvakottai, Pudukkottai and Thiruvarankulam of Pudukkottai district in Tamil Nadu (Table 1).
Table 1: Survey and collection of disease images from rice fallow black gram cultivated areas of Tamil Nadu
<table><tr><td>S.
No.</td><td>Survey
Conducted
Villages</td><td>Geo
Co-
ordinates</td><td>Name of the
Blocks</td><td>Name of the
District</td><td>Cultivars</td></tr><tr><td>1.</td><td>Vannarapatti</td><td>10.52;78.96</td><td>Pudukkottai</td><td></td><td></td></tr><tr><td>2.</td><td>Vannarapatti</td><td>10.52;78.96</td><td>Pudukkottai</td><td></td><td></td></tr><tr><td>3.</td><td>Adanakottai</td><td>10.53;78.96</td><td>Pudukkottai</td><td></td><td></td></tr><tr><td>4.</td><td>Adanakottai</td><td>10.53;78.97</td><td>Pudukkottai</td><td></td><td></td></tr><tr><td>5.</td><td>S.Solagampatti</td><td>10.59;79.02</td><td>Gandarvakottai</td><td></td><td></td></tr><tr><td>6.</td><td>S.Solagampatti</td><td>10.59;79.01</td><td>Gandarvakottai</td><td></td><td></td></tr><tr><td>7.</td><td>S.Solagampatti</td><td>10.59;79.01</td><td>Gandarvakottai</td><td></td><td></td></tr><tr><td>8.</td><td>S.Solagampatti</td><td>10.59;79.01</td><td>Gandarvakottai</td><td></td><td></td></tr><tr><td>9.</td><td>Puthunagar</td><td>10.62;79.04</td><td>Gandarvakottai</td><td></td><td></td></tr><tr><td>10.</td><td>Puthunagar</td><td>10.62;79.04</td><td>Gandarvakottai</td><td></td><td></td></tr><tr><td>11.</td><td>Puthunagar</td><td>10.43;78.98</td><td>Gandarvakottai</td><td></td><td></td></tr><tr><td>12.</td><td>Puthunagar</td><td>10.43;78.97</td><td>Gandarvakottai</td><td rowspan="2">Pudukkottai</td><td rowspan="2">VBN8, VBN10 & VBN11</td></tr><tr><td>13.</td><td>Puthunagar</td><td>10.42;78.97</td><td>Gandarvakottai</td></tr><tr><td>14.</td><td>Vellalaviduthi</td><td>10.51; 79.07</td><td>Gandarvakottai</td></tr><tr><td>15.</td><td>Mankkottai</td><td>10.35; 78.95</td><td>Thiruvarankulam</td></tr><tr><td>16.</td><td>Mankkottai</td><td>10.35; 78.91</td><td>Thiruvarankulam</td></tr><tr><td>17.</td><td>Kothakottai</td><td>10.34;78.91</td><td>Thiruvarankulam</td></tr><tr><td>18.</td><td>Dhatchinapura m</td><td>10.34; 78.91</td><td>Thiruvarankulam</td></tr><tr><td>19.</td><td>Pappanpatti</td><td>10.37; 78.91</td><td>Thiruvarankulam</td></tr><tr><td>20.</td><td>NPRC, Vamabn</td><td>10.36; 78.92</td><td>Thiruvarankulam</td></tr><tr><td>21.</td><td>Ambalapuram</td><td>10.19; 79.13</td><td>Aranthangi</td></tr><tr><td>22.</td><td>Sengarai,</td><td>10.11; 78.93</td><td>Aranthangi</td></tr><tr><td>23.</td><td>M. S. K. Veedu</td><td>10.22; 78.90</td><td>Aranthangi</td></tr><tr><td>24.</td><td>Kunnakkurumb i,</td><td>10.21; 78.97</td><td>Aranthangi</td></tr><tr><td>25.</td><td>Ramasamipura m</td><td>10.23; 79.13</td><td>Aranthangi</td></tr><tr><td>26.</td><td>Mangalanadu</td><td>10.20; 79.14</td><td>Aranthangi</td></tr><tr><td>27.</td><td>Kallaakottai</td><td>10.50; 79.08</td><td>Aranthangi</td></tr></table>
The diseases (anthracnose, powdery mildew, leaf crinkle, yellow mosaic) infected and additionally healthy leaf sample images were captured by a camera (SONY Alpha ILCE-6100Y APS-C) in cultivars of VBN8, VBN10, and VBN11 during sunlight of morning 6.00 to 10.00 am. Totally, 27376 images were collected at 24.2 Mega Pixels (Each one), which comprised disease images such as anthracnose (4225 Nos.), leaf crinkle (6354 Nos.), powdery mildew (6120 Nos.), yellow mosaic (4211 Nos.) and healthy (6466 Nos.). Furthermore, these images were classified by the ratio (80:20) several training sets and several test sets on each and every disease and accompanied by typical symptoms [19], and were described in Table 2.
Table 2: Classification of Disease Images by Application of Using Sets and Symptomatology
<table><tr><td rowspan="2">Images Classificatio n (Diseases)</td><td colspan="2">No. of Using Sets of Images</td><td rowspan="2">Total No. of using Sets</td><td rowspan="2">Typical Symptoms</td></tr><tr><td>Training Set</td><td>Test Set</td></tr><tr><td>Anthracnose</td><td>3,380</td><td>845</td><td>4,225</td><td>Circular, black, sunken spots with a dark center and bright red-orange margins on leaves [20].</td></tr><tr><td>Leaf crinkle</td><td>5,084</td><td>1,270</td><td>6,354</td><td>The youngest leaves as chlorosis around some lateral veins and branches near the margin. The leaves show curling of the margin downwards.Some of the leaves show twisting. The veins show reddish-brown discoloration on the under surface which also extends to the petiole [21].</td></tr><tr><td>Powdery Mildew</td><td>4,896</td><td>1,224</td><td>6,120</td><td>White powdery patches appear on leaves and other green parts which later become dull-colored. These patches gradually increase in size and become circular covering the lower surface completely [22].</td></tr><tr><td>Yellow Mosaic</td><td>3,201</td><td>1,010</td><td>4211</td><td>Initially, mild scattered yellow spots appear on young leaves and show irregular yellow and green patches alternating with each other. Spots gradually increase in size and ultimately some leaves turn completely yellow. Infected leaves also show necrotic symptoms [23].</td></tr><tr><td>Healthy</td><td>4,801</td><td>1,665</td><td>6466</td><td>Uninfected</td></tr><tr><td>Total No. of images used</td><td>21,362</td><td>6,014</td><td>27,376</td><td></td></tr></table>
### b) Image Pre-Processing
In our experiments, image pre-processing and CNN algorithms, as shown in Figure 1, were implemented using Anaconda 3(Python 3) and the Keras library. The experimental hardware environment includes a Mac Pro with $1.4\mathrm{GHz}$ Quad-Core Intel Core i5. The symptoms-based classified images were further used in this pre-processing. The images were preprocessed using Python programming language with image width, height $150\times 150$ with images $256\times 256$ pixels. Images were trained using the inception V3 model. Images were divided into $80\%$ training and $20\%$ testing. In each step, there were different numbers of batches, and each batch contained about 32 images. Data augmentation such as flip, rotation, and zoom were used to create an anomaly dataset for training the model and to detect anomalies. The background noise for images was zoomed out to remove the unnecessary features and to overcome the dropouts. Further, the convolution layers were used to make clarification and diagnosis of diseases with similarity scores and dissimilarities.
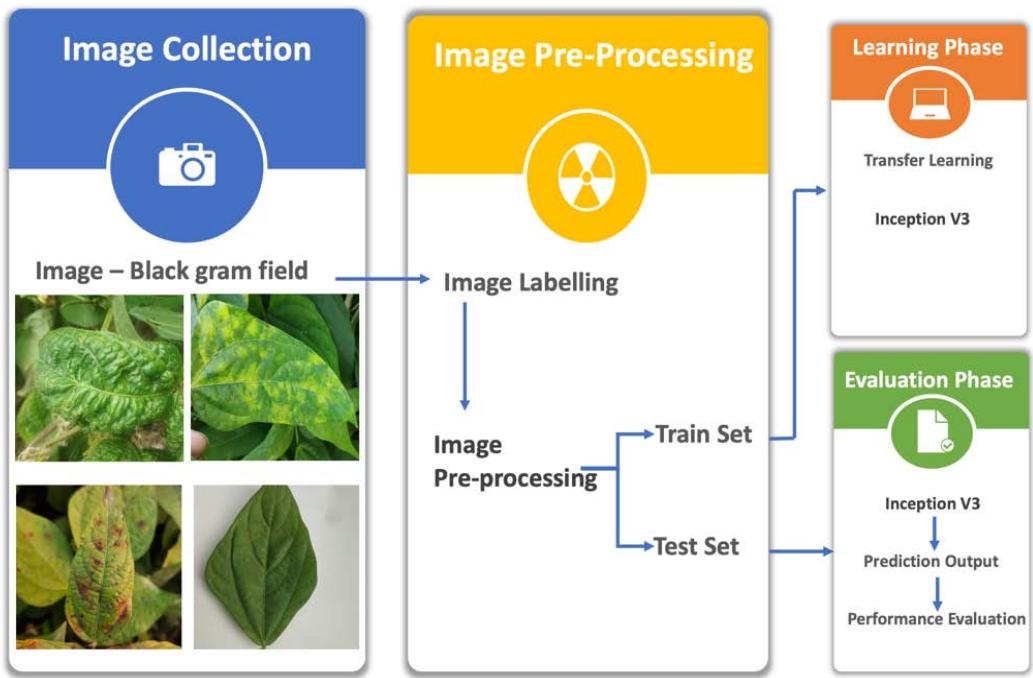 Figure 1: The Diagram to Identify Black Gram Disease
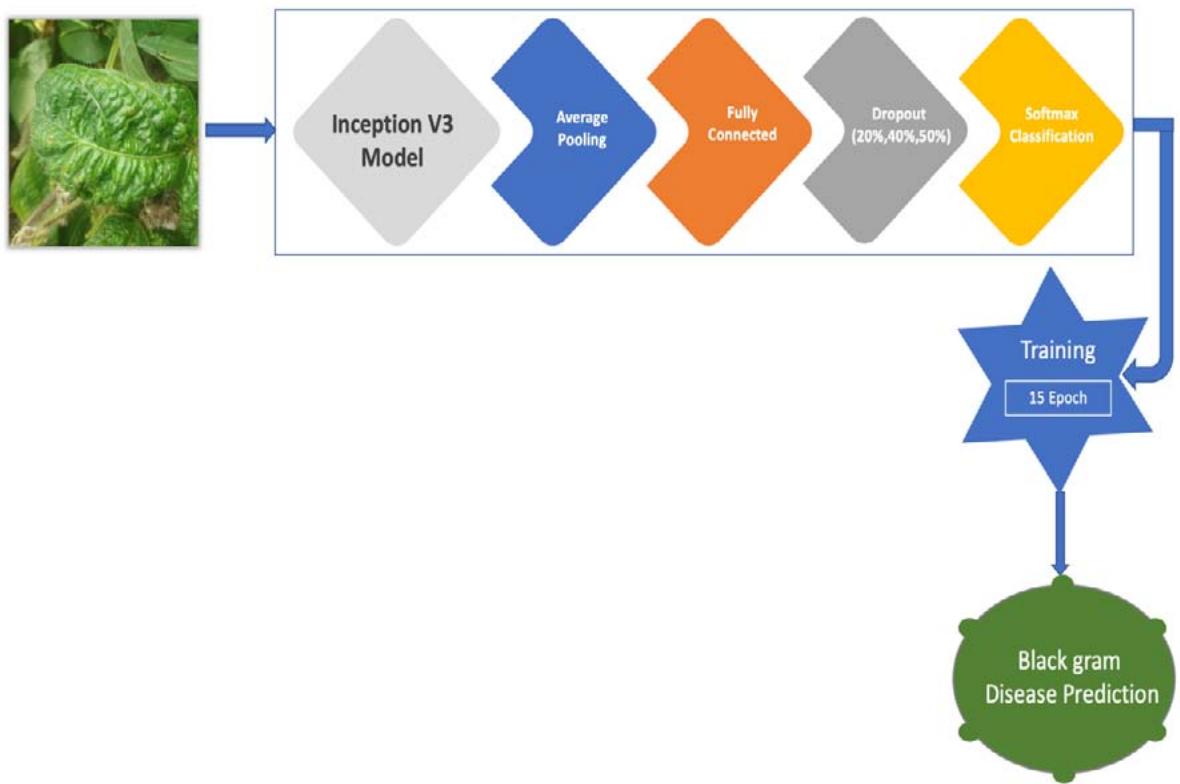 Figure 2: CNN Architecture for the Pre-Trained Model for Black Gram Disease Diagnosis
### c) Application of Convolution Layers
In this layer, the filters or kernels were used to extract the features from the black gram images by moving the kernel over the images, and with the features, feature maps were created. Backpropagation techniques were used with filters and weights. In this layer, the number of parameters was reduced to maintain the network efficiency.
## i. Batch Normalization Layers
In this layer data were broken down into mini batches during normalization, to reduce the complexity of the system. In this layer the speed-up process was initiated for training deep neural network features $z$ is remeasured with the equation
$$
Z_{norm}^{(i)} = \frac{z^{(i)} - \mu_B}{\sqrt{\sigma_{B - \varepsilon}^2}} \tag{1}
$$
Where
- = very small positive number
- = A verage of mini batch mean
- = Mini batch variance
The value of $z$ in the first layer is calculated according to the equation
$$
z = \omega x + b
$$
where
$x =$ the value of input features
$$
b = bias
$$
To scale and shift the normalized input $\gamma$ and $\beta$ are added, learning using the network parameters through equation
$$
\widetilde{z^{(i)}} = \gamma z_{norm}^{(i)} + \beta
$$
## ii. Activation Layers
In this layer, the activation function was used to speed up the arithmetic operation without exploiting the problems. Rectified Linear Unit (ReLU) was used as the basic activation function $\mathbf{z}$ using a mathematical model
$$
\mathbf{a} = \max\left(0; \tilde{\mathbf{\Sigma}}_z\right) \tag{4}
$$
The final activation function was used as the output for the last dense layer. The prediction probabilities were 0, 1, 2, 3, 4 for multiple classifications. The softmax function was used for the decimal probabilities of each class. It was calculated from output i with the equation
$$
a ^ {(i)} = \frac {e ^ {z ^ {(i)}}}{\sum_ {i} ^ {m} e ^ {z ^ {(i)}}} \quad \text {f o r i = 1 , 2 , 3 . . . m .} \tag {5}
$$
where $z(i) =$ the output of the i dimension
$$
a(i) = probability related to i class
$$
$\mathsf{m} =$ number of dimensions with respect to the number of classes
The class with the highest probability, while predicting by the method was as described below
$$
\begin{array}{l} \widehat {y} _ {t} = \max a ^ {(i)} \\i \varepsilon [ l, m ] \tag {6} \\\end{array}
$$
## iii. Pooling Layers
In these layers, the dimension of feature maps was reduced, selecting the important features created by the convolution layers. There were two poolings: average and maximum pooling. In our model, average pooling, which takes the average value of each feature map, was used.
## iv. Fully Connected Layers
In these layers, there were neurons and the last layer of the neural network. With the input, the final output of the pooling or convolution layers is turned into
$$
L _ {\text{cross} \_ \text{entropy}} (\hat{y}, \mathrm{y}) = - \sum_ {j = 0} ^ {K} y _ {j} \log (\hat{y} _ {j})
$$
single vector by means of the flattened layer. The ediction was done using weights, and the final obabilities were given by the dense layer. The number parts in the last dense layer which corresponds to the number of classes.
## v. Dropout layer
This layer was used to regularize the neural network and to avoid overfitting by deleting the portion of incoming neurons and their connection.
### d) Functions and Optimizations
## i. Loss function
This was also called as cost function. The cross-entropy was to find the loss between actual output $y$ and the expected output $\hat{y}$. Cross entropy for multiple classifications was calculated using the equation
$$
f o r j = 1, 2, 3 \dots k \tag {7}
$$
$$
k= number of classes
$$
## ii. Optimization Function
The optimization function was used to reduce the loss function; the Adam optimizer was used in this model. Weights were learned adaptively by Adam Optimizer.
### e) Assessment and Evaluation
## i. Application of Inception V3 models
In the research paper, Inception V3 was used for pre-training the models. Inception V3 was an additional design for CNN developed by Google. Inception starts with a sparse structure, increases the network depth and width, and clusters the spare data into a dense structure to enhance the model accuracy [26].
Transfer the learning of plant pathology data to the Inception V3 model pre-trained on ImageNet data to speed up the training process and to improve the model performance; the Inception V3 model has 94 convolution layers, 14 pooling, and dense layers [25]. Transfer of learning, pre-trained CNN model includes feature extraction and fine-tuning to achieve the best results. In this paper, feature extraction was first used to train the new classifier, and the model was fine-tuned. Inception V3 was trained using training parameters (Table 3).
Table 3: Parameters value to build black gram disease diagnosis for CNN models
<table><tr><td>Parameter</td><td>Value</td></tr><tr><td>Batch Size</td><td>32</td></tr><tr><td>Dropout</td><td>20%, 40%, and 50%</td></tr><tr><td>Activation function</td><td>ReLU, SoftMax</td></tr><tr><td>Optimizer</td><td>Adam</td></tr><tr><td>CNN Training</td><td></td></tr><tr><td>Epoch</td><td>15</td></tr><tr><td>Epoch Learning Rate</td><td>0.001</td></tr></table>
Optimization level for Batch size: $>30$; Dropout: 20, 40 & 50% (Study carried out in this experiment at V3 Inception Model) and Epoch 15 is optimized one (Constant) In the research paper, average pooling, dense layer, and SoftMax were added as the activation function for the last layer. The dropout rate was used to train the model. The next step was determining the learning rate, and Adam was used as an optimizer. The
epochs specified were 15 for the training set. The Inception V3 model consists of 21,768,352 trainable parameters during CNN training. The Inception V3 model used to diagnose black gram diseases is summarized in Table 4.
Table 4: Summary of black gram disease diagnosis using Inception V3 models
<table><tr><td>Type of Layer</td><td>Output Shape</td><td>Parameters</td></tr><tr><td>Input</td><td>(224, 224, 3)</td><td>0</td></tr><tr><td>Sequential</td><td>(224, 224, 3)</td><td>0</td></tr><tr><td>Functional (Inception V3)</td><td>(5, 5, 192)</td><td>393216</td></tr><tr><td>Average Pooling</td><td>(5, 5, 1280)</td><td>0</td></tr><tr><td>Dropout</td><td>(5, 5, 1280)</td><td>0</td></tr><tr><td>Dense</td><td>(0,3)</td><td>512</td></tr><tr><td>Total parameters</td><td>21,802,784</td><td></td></tr><tr><td>Trainable parameters</td><td>21,768,352</td><td></td></tr><tr><td>Non-trainable parameters</td><td>34,432</td><td></td></tr></table>
CNN has numerous parameters, so there was a possibility of overfitting issues, which can be solved through the dropout procedure. Dropout was a method of randomly disconnecting that can separate connections across different nodes and had a 1-p dropout probability. The dropout layer increases the algorithm's robustness while reducing the number of model parameters. The random inactivation layer enhances the robustness of the network structure and helps to prevent overfitting in the model [26]. Dropout has recently been utilized to reduce overfitting in deep neural networks. Dropout disabled certain neurons in each layer during each epoch, and the remaining neurons were used for forward and backward propagations. As a result, the active neurons were motivated to extract the needed features independently and successfully without the assistance of inactive ones. In this research, dropout rates were 20 percent, 40 percent and 50 percent were studied. Assessment indicators used for the classification of disease diagnosis were accuracy and confusion matrix for comparing models when training and testing the image dataset used. The confusion matrix is a table that shows the classification model performance on the test image set by matching the actual output and expected outputs to identify the correct disease diagnosis. The accuracy is the percentage of valid predictions from the forecast made as per the equation listed below and expressed in percentage.
$$
\text{Accuracy} = \frac{\text{Numberofcorrectprediction}}{\text{Totalnumberofprediction}} = \frac{\mathrm{T P} + \mathrm{T N}}{\mathrm{T P} + \mathrm{T N} + \mathrm{F P} + \mathrm{F N}} \tag{8}
$$
$$
where TP = True Positive
$$
$$
TN = True Negative
$$
$$
FP = False Positive, and
$$
$$
FN = False Negative
$$
## III. EXPERIMENTAL RESULTS
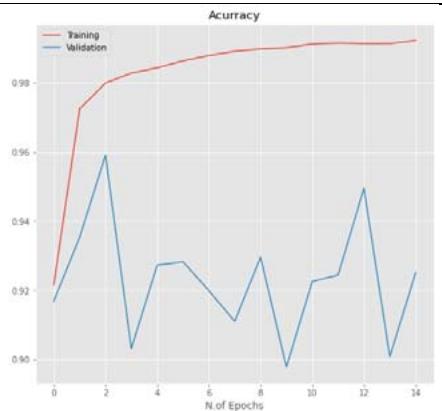
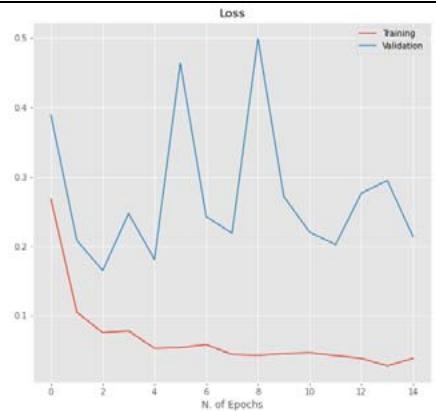
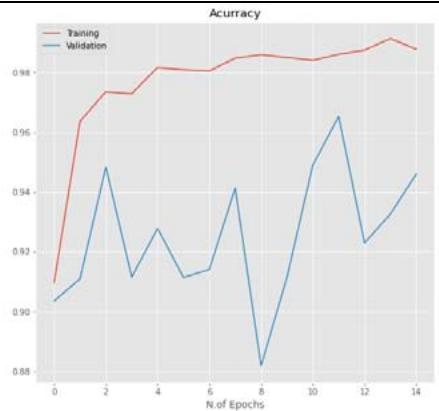
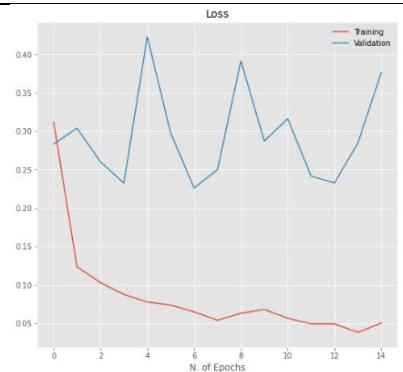
CNN model mentioned in Section 2.5 is trained using the parameters in Table 3 with dropout ratios. The model was compared with accuracy and loss while training and testing, as listed in Table 5. From the table, Inception V3's 20 percent dropout rate was the best model when compared with the 40 percent. The percentage the accuracy of the 20 percent dropout was 99.22 percent, and the loss was 0.0249. The actual and predicted result using CNN is shown in Figure 3. The confusion matrix for the model used by different dropouts is depicted in Figure 4. The changes both in the accuracy and loss of the models used in 20 percent, 40 percent, and 50 percent dropouts were shown in Figure 5. Furthermore, the results suggested that models used in the various study dropouts had an excellent ability to discriminate against black gram diseases. Therefore, based on this empirical analysis, we concluded that the models used in different study dropouts were effective in identifying black gram diseases. This study can be extended for the diagnosis of other plant diseases.
 Actual: Healthy Predicted:Leaf Crinkle
 Actual: Healthy Predicted: Healthy
 Actual: Yellow Mosaic Predicted: Yellow Mosaic
 Actual: Healthy Predicted:Healthy

 Actual: Powdery Mildew Predicted: Powdery Mildew
 Actual: Healthy Predicted: Healthy
 Actual: Healthy Predicted: Healthy

 Actual: Yellow Mosaic Predicted: Yellow Mosaic Figure 3: The actual and predicted result using CNN
Table 5: Accuracy and loss for the training, and testing phases of the Inception V3 model
<table><tr><td>Drop Out (%)</td><td colspan="2">Train</td><td colspan="2">Test</td></tr><tr><td></td><td>Accuracy (%)</td><td>Loss</td><td>Accuracy (%)</td><td>Loss</td></tr><tr><td>20</td><td>99.22</td><td>0.0249</td><td>92.51</td><td>0.2491</td></tr><tr><td>40</td><td>98.77</td><td>0.0384</td><td>94.60</td><td>0.2132</td></tr><tr><td>50</td><td>98.27</td><td>0.0506</td><td>90.32</td><td>0.3762</td></tr></table>
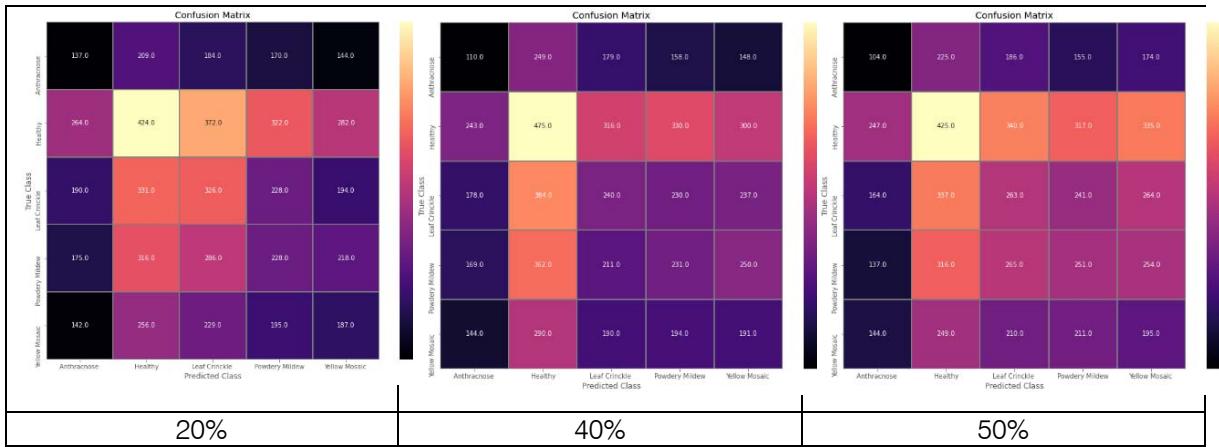 Figure 4: Confusion matrix of black gram leaf disease for Inception V3 model with $20\%$, $40\%$, and $50\%$ dropout values
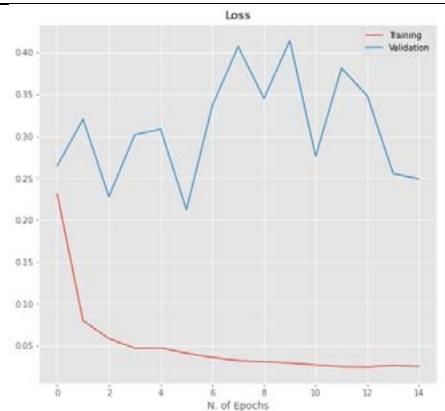
 20%
 40%
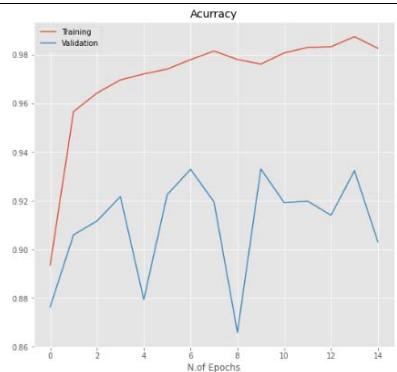 50% Figure 5: The relationship between accuracy and loss with the number of epochs for the image dataset for the Inception V3 model with $20\%$, $40\%$, and $50\%$ dropout values.
## IV. DISCUSSION
The Inception V3 model was employed in this study to diagnose foliar diseases in black gram. Relevant literature was reviewed, and the performance of the Inception V3 model was compared with existing methods. The resulting accuracy obtained is highlighted in Table 6. Deep learning models such as DenseNet, ResNet, and GoogleNet were used to detect tomato diseases [29-33]. The results obtained using these models ranged between 91 percent to 97 percent accuracy. From the proposed model, the obtained accuracy was higher at 99.22 percent. Hence, it was evident that this model had significant improvement compared to other models, as shown in Table 6.
Table 6: Performance of the proposed model with other existing models employed
<table><tr><td>S. No.</td><td>Author (s)</td><td>Method</td><td>Image Number</td><td>Image Source</td><td>Accuracy (%)</td></tr><tr><td>1.</td><td>Agarwal et al. (2020) [25]</td><td>CNN network</td><td>17,500</td><td>Plant Village</td><td>91.20</td></tr><tr><td>2.</td><td>Prajwala Tm et al. (2018) [26]</td><td>LeNet based CNN</td><td>18,160</td><td>Plant Village</td><td>95</td></tr><tr><td>3.</td><td>Widiyanto et al. (2019) [24]</td><td>CNN model</td><td>5000</td><td>Plant Village</td><td>96.60</td></tr><tr><td>4.</td><td>Keke Zhang et al. (2018) [23]</td><td>ResNet</td><td>5550</td><td>Plant Village</td><td>97.28</td></tr><tr><td>5.</td><td>Proposed model</td><td>Inception V3</td><td>27376</td><td>TNAU field</td><td>99.22</td></tr></table>
## V. CONCLUSIONS
Detection and diagnosis of plant diseases at their earlier stages play a superior role in disease management in crop protection. Keeping this strategy, this work was framed and carried out using deep learning algorithms to diagnose foliar diseases in black gram. This work entirely depends on deep neural networks and their efficacy in image processing, assessment of accuracy, and evaluation. Inception V3 models were used to retrain the transfer learning. The last layer of the models is removed and replaced with average pooling, fully connected, and softmax, respectively. Different dropout rates were used, such as 20 percent, 40 percent, and 50 percent. The experimental results showed that a high degree of accuracy was obtained in diagnosing diseases in black gram using deep learning algorithms. It was reported that a high degree of accuracy of 99.22 percent and a reported loss of 0.0249 percent were obtained in this study by using deep learning algorithms. Further, the image dataset can be expanded for more accurate and high-end results. Under field conditions, manual visualization can lead to errors in plant disease diagnosis due to abiotic factors such as light intensity, weather, and manual handling errors. However, with this approach, these factors have no effect from the field to the laboratory. So, future investigation of this research will focus on diagnosing other plant diseases.
### ACKNOWLEDGMENT
I want to express my deepest gratitude to the authors of Saeed, A.; Abdel-Aziz, A.A.; Mossad, A.; Abdelhamid, M.A.; Alkhaled, A.Y.; Mayhoub, M for their valuable research inputs in the paper Smart Detection of Tomato Leaf Diseases Using Transfer Learning-Based Convolutional Neural Networks.
Generating HTML Viewer...
References
32 Cites in Article
I Buja,E Sabella,A Monteduro,M Chiriaco,L De Bellis,A Luvisi,G Maruccio (2021). Advances in plant disease detection and monitoring: from traditional assays to in-field diagnostics.
Ning Zhang,Guijun Yang,Yuchun Pan,Xiaodong Yang,Liping Chen,Chunjiang Zhao (2020). A Review of Advanced Technologies and Development for Hyperspectral-Based Plant Disease Detection in the Past Three Decades.
Luis Rubio,Luis Galipienso,Inmaculada Ferriol (2020). Detection of Plant Viruses and Disease Management: Relevance of Genetic Diversity and Evolution.
Yi Fang,Ramaraja Ramasamy (2015). Current and Prospective Methods for Plant Disease Detection.
Houda Orchi,Mohamed Sadik,Mohammed Khaldoun,Essaid Sabir (2023). Automation of Crop Disease Detection through Conventional Machine Learning and Deep Transfer Learning Approaches.
Maha Altalak,Mohammad Ammad Uddin,Amal Alajmi,Alwaseemah Rizg (2022). Smart Agriculture Applications Using Deep Learning Technologies: A Survey.
M Anne-Katrin (2016). Plant disease detection by imaging sensors-parallels and specific demands for precision agriculture and plant phenotyping.
N Mamat,M Othman,R Abdoulghafor,S Belhaouari,N Mamat,S Hussein (2022). Advanced technology in agriculture industry by implementing image annotation technique and deep learning approach: a review.
Rikiya Yamashita,Mizuho Nishio,Richard Do,Kaori Togashi (2018). Convolutional neural networks: an overview and application in radiology.
J Boulent,S Foucher,J Theau,P St-Charles (2019). Convolutional neural networks for the automatic identification of plant diseases.
J Andrew,J Eunice,D Popescu,M Kalpana Chowdary,J Hemnath (2022). Deep learning-based leaf disease detection in crops using images for agricultural applications.
Srinivas Talasila,Kirti Rawal,Gaurav Sethi,Sanjay Mss (2022). Black gram Plant Leaf Disease (BPLD) dataset for recognition and classification of diseases using computer-vision algorithms.
A Ahmad,D Saraswat,A El Gamal (2023). A survey on using deep learning techniques for plant disease diagnosis and recommendations for development of appropriate tools.
S Daryanto,L Wang,J Pierre-Andre (2015). Global synthesis of drought effects on food legume production.
R Ajaykumar,P Prabakaran,K Sivasabari (2022). Growth and Yield Performance of Black Gram (Vigna mungo L.) under Malabar Neem (Melia dubia) Plantations in Western Zone of Tamil Nadu.
Pavel Nazarov,Dmitry Baleev,Maria Ivanova,Luybov Sokolova,Marina Karakozova (2020). Infectious plant diseases: etiology, current status, problems and prospects in plant protection.
B Punam,P Gole (2021). Plant disease detection using hybrid model based on convolutional autoencoder and convolutional neural network.
A Reshmi,P Prasidhan (2022). Leaf disease detection using CNN.
S Aggarwal,B Mali,P Rawal (2015). MANAGEMENT OF ANTHRACNOSE OF FIELD BEAN CAUSED BY COLLETOTRICHUM LINDEMUTHIANUM THROUGH DIFFERENT FUNGICIDES AND BIOAGENTS.
Adhimoolam Karthikeyan,Manoharan Akilan,Santhi Samyuktha,Gunasekaran Ariharasutharsan,V Shobhana,Kannan Veni,Murugesan Tamilzharasi,Krishnan Keerthivarman,Manickam Sudha,Muthaiyan Pandiyan,Natesan Senthil (2022). Untangling the Physio-Chemical and Transcriptional Changes of Black Gram Cultivars After Infection With Urdbean Leaf Crinkle Virus.
M Jayasekhar,E Ebenezar (2016). Management of powdery mildew of black gram (<italic>Vigna mungo</italic>) caused by <italic>Erysiphe polygoni</italic>.
S Kothandaraman,D Alice,V Malathy (2016). Seed-borne nature of a begomovirus.
Ashraf Darwish,Dalia Ezzat,Aboul Hassanien (2020). An optimized model based on convolutional neural networks and orthogonal learning particle swarm optimization algorithm for plant diseases diagnosis.
J Chen,J Chen,D Zhang,Y Sun,Y Nanehkaran (2020). Using deep transfer learning for image-based plant disease identification.
Al Husaini,M Habaebi,M Gunawan,T Islam,M Elsheikh,E Suliman,F (2022). Thermal-based early breast cancer detection using inception V3, inception V4 and modified inception MV4.
C Szegedy,V Vanhoucke,S Ioffe,J Shlens,Z Wojna (2015). Rethinking the inception architecture for computer vision.
Nauman Munir,Hak-Joon Kim,Sung-Jin Song,Sung-Sik Kang (2018). Investigation of deep neural network with drop out for ultrasonic flaw classification in weldments.
Keke Zhang,Qiufeng Wu,Anwang Liu,Xiangyan Meng (2018). Can Deep Learning Identify Tomato Leaf Disease?.
Sigit Widiyanto,Rizqy Fitrianto,Dini Wardani (2019). Implementation of Convolutional Neural Network Method for Classification of Diseases in Tomato Leaves.
M Kalpana,L Karthiba,K Senguttuvan,R Parimalarangan (2023). Diagnosis of Major Foliar Diseases in Black gram (Vigna mungo L.) using Convolution Neural Network (CNN).
No ethics committee approval was required for this article type.
Data Availability
Not applicable for this article.
How to Cite This Article
Dr. Selvanayaki Kolandapalayam Shanmugam. 2026. \u201cAdvancing Black Gram (Vigna Mungo L.) Disease Diagnosis Through Deep Learning Techniques\u201d. Global Journal of Computer Science and Technology - G: Interdisciplinary GJCST-G Volume 24 (GJCST Volume 24 Issue G1): .
Explore published articles in an immersive Augmented Reality environment. Our platform converts research papers into interactive 3D books, allowing readers to view and interact with content using AR and VR compatible devices.
Your published article is automatically converted into a realistic 3D book. Flip through pages and read research papers in a more engaging and interactive format.
Interestingly, in sustainable crop protection, disease diagnosis, and management are crucial in sustainable crop production. It plays a captious role in rain-fed pulses because the occurrence of season of cropping, cultivation after main crop, availability of soil moisture in poor conditions, consecutively following the same cultivars are acting a predominant role in disease diagnosis approaches and confirmation. Under these situations, occurrence of the manual errors (or) mis find faults resulting in complete drawbacks to disease diagnosis and management for farmers and scientists worldwide. Keeping this background, applying deep learning techniques is most helpful in diagnosing plant diseases silently and superiorly. Deep learning techniques were carried out in this study to diagnose foliar diseases in black gram such as anthracnose, leaf crinkle, powdery mildew, and yellow mosaic that causes a severe yield loss (>50%) silently accompanied by green biomass.
Our website is actively being updated, and changes may occur frequently. Please clear your browser cache if needed. For feedback or error reporting, please email [email protected]
Thank you for connecting with us. We will respond to you shortly.