## I. INTRODUCTION
Cancers are now the leading cause of premature death in 127 countries. There is a trend that cancer may surpass cerebral vascular disease (CVD) as the leading cause of sudden death in most countries (Bray et al., 2021). According to the International Agency for Research on Cancer, 19.3 million new cancer cases and almost 10.0 million cancer deaths occurred in 2020. The global cancer burden is expected to be 28.4 million cases in 2040, a $47\%$ rise from 2020 (Sung et al., 2021). Cancer is essentially a disease of the genome, which evolves and progresses with accumulations of somatic mutations, including copy-number alterations, structural variants, and epigenomic alterations (Nakagawa et al., 2018; Stratton et al., 2009). Thus, a pan-cancer analysis of cancer-associated genes will help understand their roles in cancer development.
We have identified abnormal expression of Peptidylprolyl isomerase B (PPIB) in various cancers through preliminary database analysis. PPIB is a member of the cyclosporine-binding protein and is mainly located within the endoplasmic reticulum. It is associated with the secretory pathway and released in biological fluids. This protein can bind to cells derived from T- and B-lymphocytes and may regulate the immunosuppressive drug cyclosporine A-mediated immunosuppression as a cell receptor protein. The molecular function of PPIB is to act as a peptidylprolyl anti-cis isomerase (PPIase), regulating the protein conformation of its substrate through propyl cis-trans isomerization in the endoplasmic reticulum lumen and nucleus, and participating in protein folding, secretion, and post-translational modification processes. PPIB could catalyze the cis-trans isomerization of xaa-proline bonds, a rate-limiting step in protein folding, which is required for proteome homeostasis (Hasel KW et al., 1991 Jul; Peddada LB et al., 1992).
There have been some relevant studies on the role of PPIB in cancer. A recent study found that UTMD-mediated miR-206 regulates the expression of apoptosis, migration, and invasion-related proteins by targeting PPIB, inhibiting the migration and invasion of hepatocellular cancer (LIHC) cells, and enabling cell apoptosis (Wu H et al., 2020 Jan-Dec). PPIB can enhance the JAK2/SAT3 signaling pathway in gastric cancer cells, thereby promoting the proliferation of cancer cells (Li T et al., 2017 Mar 1). Furthermore, PPIB can serve as a biomarker in the diagnosis and treatment of cancer. In ovarian cancer (OV) and head and neck squamous cell carcinoma (HNSC), PPIB is a potential prognostic marker (Pan X & X., 2020 Oct 15; Xin et al., 2021 Mar 29). PPIB induces chemotherapy resistance by degrading wild-type p53 through its interaction with MDM2, which may serve as a predictive biomarker for chemotherapy resistance in the treatment of colorectal cancer (Choi et al., 2018 Sep). In addition, another study found that PPIB is also a novel indicator for evaluating lymph node metastasis in colorectal cancer (Yue et al., 2009 Oct).
This study focused on PPIB and utilized bioinformatics-related software and online databases to comprehensively analyze the differences and specific molecular mechanisms of PPIB in pan-cancer, providing a comprehensive understanding and sufficient basis for whether PPIB has the potential for therapeutic targets and detection markers in human cancers.
## II. MATERIALS AND METHODS
### a) Research data sources
The data for this study is sourced from TCGA and GTEx databases. We analyzed the expression level of PPIB and its correlation with CAFs using the TIMER2.0 database (http://timer.comp-genomics.org/). We studied the expression level of PPIB, expression differences in pathological staging, survival analysis, and correlation analysis of co-expressed genes using the GEPIA2 database (http://gepia.cancer-pku.cn/). We explored the protein expression levels of PPIB in different tumors using the UALCAN database (http://ualcan.path.uab.edu/). We investigated the genetic variation frequency and types of PPIB in various tumors using the cBiportal database (https://www.cbiportal.org/). We dissected the correlation between PPIB and methylation using the GSCA database (https://bioinfo.life.hust.edu.cn/gsca). We analyzed the correlation between PPIB and abundance of tumor-infiltrating lymphocytes by TISIDB database (http://cis.hku.hk/tisidb/index.php). We performed protein-protein interaction network (PPI) analysis using the STRING database (https://string-db.org/). We performed KEGG pathway and GO analysis of PPIB using the DAVID database (https://david.ncifcrf.gov/). The data and analysis provided by these databases and tools provide a foundation for us to comprehensively explore the role of PPIB in cancer.
### b) Analysis Methods
## i. Expression Analysis of PPIB
We used the "Gene-DE" module in TIMER2.0 to analyze the differential expression of PPIB in different tumors and adjacent normal tissues in the TCGA database (Li et al., 2020 Jul 2). Due to the loss of normal tissue samples of some tumors in the TIMER2.0 database, we used the GEPIA2 database for analysis and supplementation of tumors with differential expression of PPIB (Tang et al., 2019 Jul 2). By using the "Pathological Stage Plot" module tool and the "CPTAC" module in the UALCAN database, we analyzed the correlation between PPIB expression and clinical staging of different cancers and the protein expression of PPIB in various cancers (Chandrashekar et al., 2017 Aug). Comparisons between groups were performed using two independent sample t tests. The difference was statistically significant at $P < 0.05$.
## ii. Survival Analysis of PPIB
We analyzed the relationship between the high/low expression status of PPIB and overall survival (OS)/disease-free survival (DFS) using the "Survival Analysis" module of the GEPIA2 database(Tang et al., 2019 Jul 2).
## iii. Genetic Variation of PPIB
We used the cBiportal database "TGCA PanCancer Atlas Study" module to analyze the genetic variation of PPIB in different tumors and display specific domain mutation information. Simultaneously explored the relationship between PPIB mutations/non-mutations and survival (Cerami et al., 2012 May).
## iv. DNA methylation analysis of PPIB
We analyzed the impact of methylation status on PPIB expression in different tumors through the "Mutation" module of the GSCA database (Liu et al., 2023 Jan 19).
## v. Analysis of the Correlation between PPIB and Tumor immune Microenvironment
We used the "Gene" module of the "Immune" module in the TIMER2.0 database to analyze the relationship between PPIB and CAFs in different tumors. Using the "Lymphocyte" module of TISIDB database, we explored the relationship between the abundance of tumor infiltrating lymphocytes and PPIB expression in various tumors (Ru et al., 2019 Oct 15).
## vi. PPIB-related gene enrichment analysis
We analyzed 50 proteins that interact with PPIB by using the STRING database "Protein by name" module (von Mering et al., 2003 Jan 1). Then, through the "Similar" module of the GEPIA2 database, we screened out the top 100 genes related to PPIB expression. We used the "Correlation Analysis" module to analyze the correlation between the top 5 genes with the highest correlation and PPIB expression. We analyzed the expression of the top 5 genes related to PPIB expression in different tumors using the "Gene Corr" module of the TIMER2.0 database. We used the DAVID database and the "OFFICIAL GENE SYN" module to screen PPIB expression-related genes from the GEPIA2 database for KEGG pathway analysis and GO analysis to provide a basis for exploring the specific molecular mechanisms and roles of PPIB in different tumors (Dennis et al., 2003).
## III. RESULTS
### a) Analysis of expression patterns of PPIB
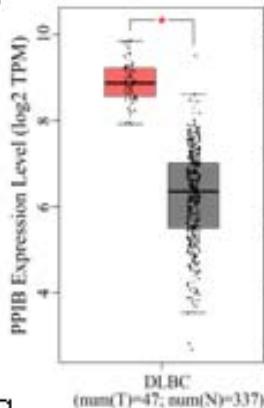
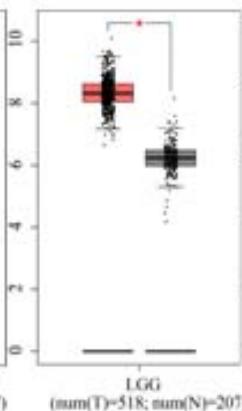
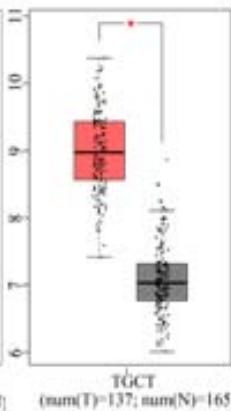
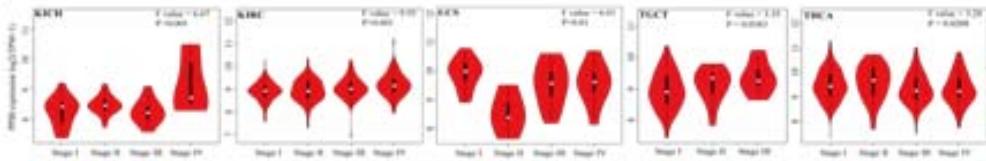
To explore the clinical significance of PPIB in tumorigenesis, we utilized the "Gene-DE" module in the TIMER2.0 database. We supplemented it with the GEPIA2 database to analyze the expression profiles of PPIB at the mRNA level in 33 types of tumors. We found that PPIB was differentially expressed in 22 types of tumor tissues compared to corresponding normal tissues, up-regulation of PPIB happened in 20 types of tumors, including bladder urothelial carcinoma (BLCA), breast invasive carcinoma (BRCA), cholangio carcinoma (CHOL), colon adenocarcinoma (COAD), esophageal carcinoma (ESCA), glioblastoma multiforme (GBM), head and neck squamous cell carcinoma (HNSC), kidney renal clear cell carcinoma(KIRC), kidney renal papillary cell carcinoma (KIRP), Liver hepatocellular carcinoma (LIHC), lung adenocarcinoma (LUAD), Lung squamous cell carcinoma (LUSC), prostate adenocarcinoma (PRAD), rectum adenocarcinoma (READ), stomach adenocarcinoma (STAD), uterine corpus endometrial carcinoma (UCEC), lymphoid neoplasm diffuse large B-cell lymphoma (DLBL), brain lower grade glioma (LGG), testicular germ cell tumors (TGCT), uterine carcinosarcoma (UCS). However, downregulation of PPIB happened in 2 types of tumors, including kidney chromophobe (KICH) and thyroid carcinoma (THCA) (Figure 1A and 1B). This indicates that PPIB may play an inhibitory role in these two types of cancer, indicating the duality of PPIB in different cancers. BRCA1 is a protein factor that protects breast tissue cells from cancer invasion. However, a recent study suggests that this protein may also have the opposite effect: in another type of cancer-neuroblastoma, BRCA1 helps stabilize the tumor (Herold et al., 2019). Additionally, we also found that the protein level of PPIB was upregulated in four kinds of tumor tissues (COAD, KIRC, LUAD, UCEC). It was shown that PPIB plays a role as a carcinogen in the above tumors. Moreover, we analyzed the expression of PPIB in different tumor pathological stages. It was suggested that the upregulation of PPIB was significantly linked to the pathological stages of KICH, KIRC, UCS, TGCT, and THCA (Figure 1C). These results indicated that PPIB may be involved in promoting the metastasis progression in these cancers.
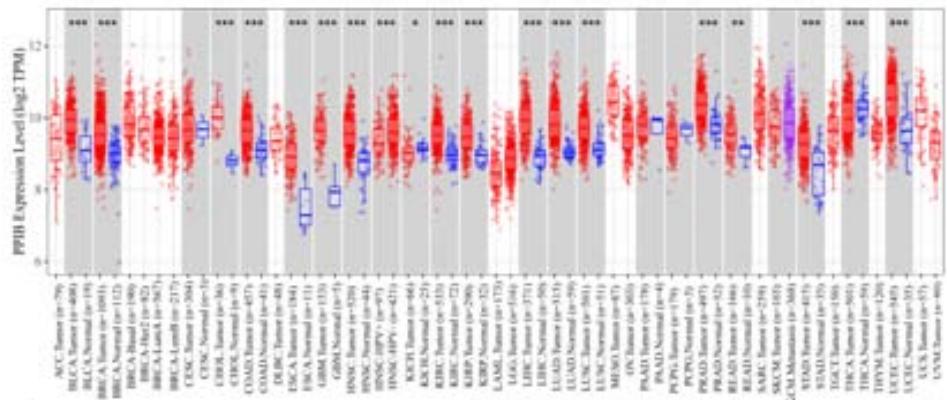 Figure 1 A
 Panel label: B.
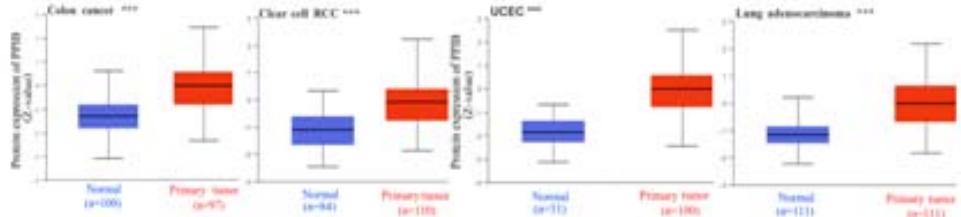


 Panel label: C.
 D Figure 1: Expression profile of PPIB in different tumors
(A) Expression of PPIB in 33 types of tumor tissues.\* $\mathsf{P} < 0.05$ $^{**}\mathrm{P} < 0.01$ $^{**}\mathrm{P} < 0.001$.(B) Expression of PPIB in DLBC, THYM, TGCT and UCS tumor tissues and corresponding normal tissues.(C) Protein expression of PPIB in primary tumor tissues and corresponding normal tissues.(D) Relationship between expression of PPIB and pathological stage.
### b) Survival Analysis of PPIB
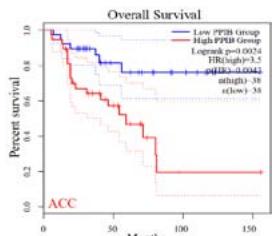
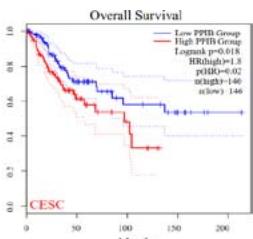
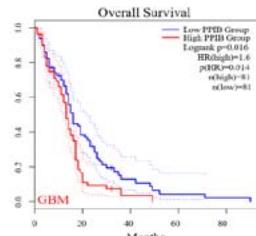
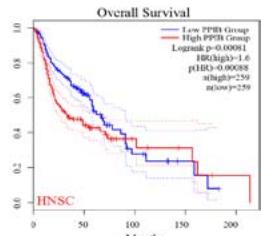
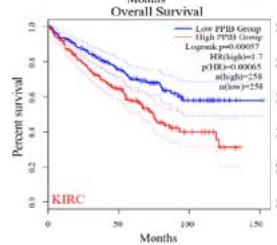
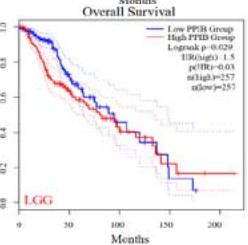
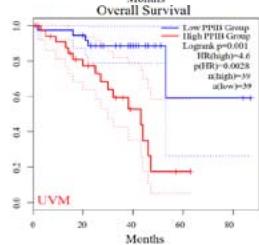
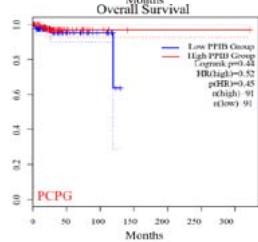
We further explored the prognostic significance of PPIB among the 33 types of tumors based on the GEPIA2 database. It was found that high expression of PPIB is negatively correlated with overall survival (OS) in ACC, CESC, GBM, HNSC, KIRC, LGG, and UVM (Figure 2A). Simultaneously, disease-free survival (DFS) analysis showed that upregulation of PPIB expression is associated with poor prognosis in ACC, CESC, CHOL,
HNSC, KIRC, LGG, and UVM (Figure 2B). These results indicated that PPIB is an independent prognostic marker of both DFS and OS in ACC, CESC, HNSC, KIRC, LGG, and UVM.
 Panel label: A.
 Panel label: B.
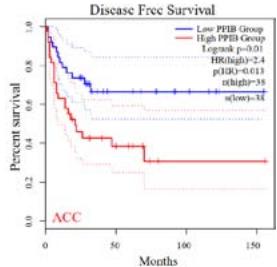
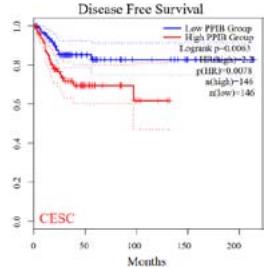
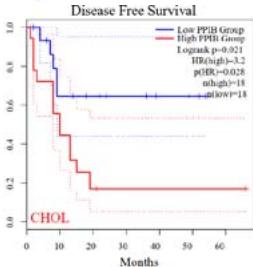
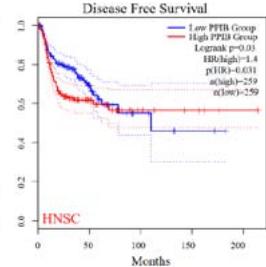
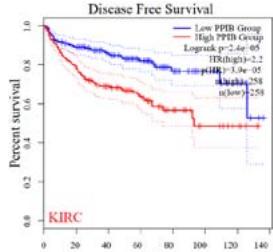
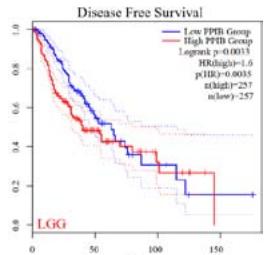
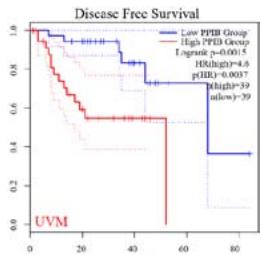 Figure 2: Survival analysis of PPIB in 33 types of tumors.
(A) Analysis of PPIB expression and overall survival in 33 types of tumors. (B)Analysis of PPIB expression and disease-free survival in 33 types of tumors.
### c) Genetic and Epigenetic Variation Analysis of PPIB
We analyzed the genetic variation of PPIB in TCGA pan-cancer using the cBiportal database. As shown in Figure 3A, the overall genetic variation frequency of PPIB is relatively low. The highest alteration frequency of PPIB happened in UCEC (2.84%), with mutations (2.46%) being the predominant. However, it is worth noting that genetic variation is mainly amplification in MESO, KICH, SARC, and PRAD. In cases of gene variation in STAD, copy number deletion (0.68%) is the leading cause. Additionally, the genetic characteristics of PPIB were not observed in tumors such as LAML, ACC, CHOL, DLBC, GBM, and PCPG (Figure 3A).
We further investigated the gene mutation types, loci, and number of cases of PPIB. As shown in Figure 3B, the highest number of cases is missense mutations among the kinds of gene mutations of PPIB. The data represented that there have been cases of missense mutations in R95C/G in BLCA, ESCA, and COAD. The arginine (R) at position 95 in the "PRO_isomerase" domain is mutated to cysteine (C) or glycine (G). In LGG, SKCM, and STAD cases, X115 spliceosomes were found on the "PRO_isomerase" domain (Figure 3B).
According to the high proportion of copy number amplification in mutations, we evaluated the relationship between PPIB expression and copy number amplification. We found a significant positive correlation between the expression levels of PPIB and copy number amplification in 20 types of cancer (FDR\<0.05), including UCEC, READ, COAD, OV, LUAD and LUSC (FDR < 0.05) (Figure 3C). Research has shown that DNA promoter methylation plays a crucial role in tumor progression (Kulis & Esteller, 2010). We evaluated the
DNA methylation patterns of PPIB based on the GSCA database. The expression level of PPIB is significantly negatively correlated with DNA methylation in 22 types of tumors (FDR $< 0.05$ ), including PRAD, BRCA, LGG, COAD, LUAD, HNSC (Figure 3D). According to the above data, DNA copy number amplification and methylation are the two underlying causes of PPIB upregulation in cancers.
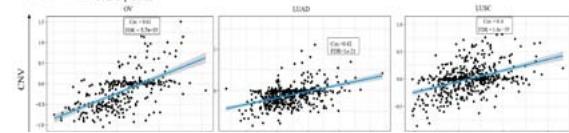
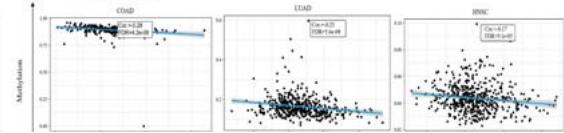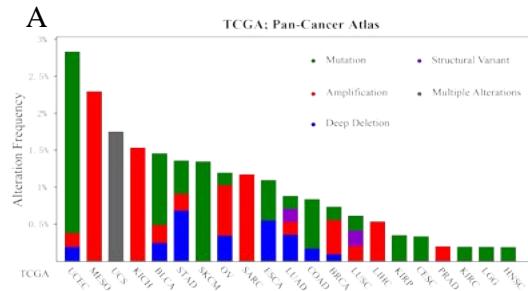

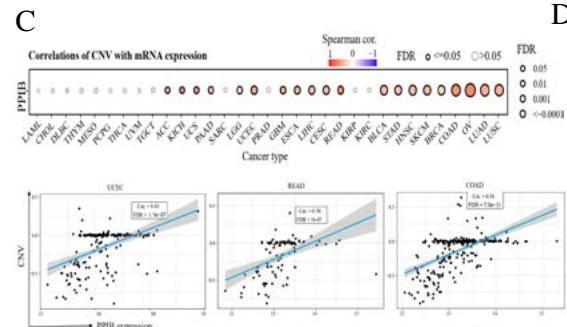


 Figure 3: Genetic variation analysis of PPIB in tumors.
- (A) Genetic variation types of PPIB in tumors. (B) Mutations in various domains of PPIB. (C) Correlation between copy number variation and PPIB expression. (D)Correlation between DNA methylation and PPIB expression.
d) Correlation analysis between PPIB and the immune microenvironment
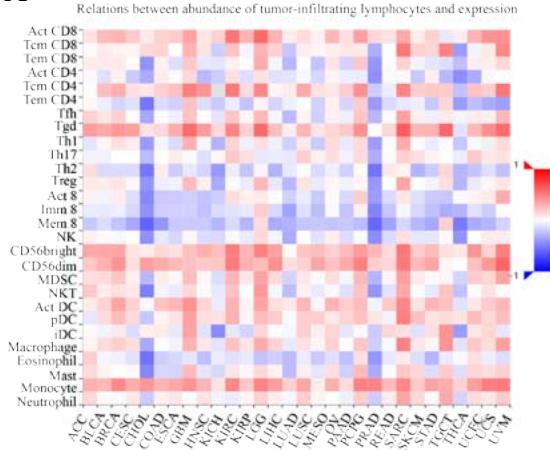
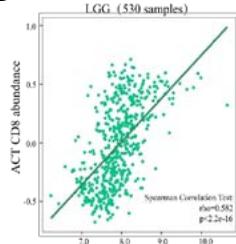
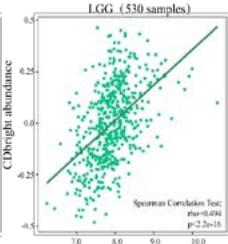
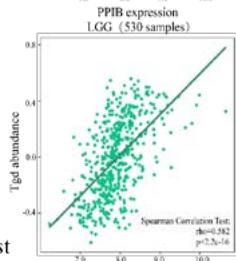
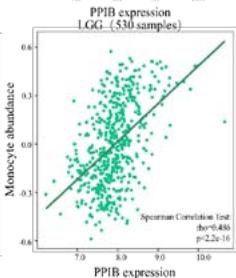
Malignant solid tumor tissue contains not only cancer cells, but also normal stromal, infiltrating immune, epithelial, and vascular cells. It has been reported that infiltrating immune cells play essential roles in regulating tumor growth, metastasis, and drug resistance (Han et al., 2020 Jun). In this study, we used the TISIDB platform to explore the correlation between PPIB expression and infiltrating immune cells. It was found that PPIB expression was positively associated with activated CD8+ T cell(Act CD8), central memory CD8+ T cell (Tcm CD8), effect memory CD8+T cells (Tem CD8), effect memory (Tcm CD4), gamma delta T cells (Tgd), CD56bright/CD56dim NK cells, myeloid suppressive cells (MDSCs) and activated dendritic cells (Act DCs). Nevertheless, we observed a statistically negative correlation between PPIB expression and activated/immature/memory B cells, natural killer cells (NK), and eosinophils (EOS) (Figure 4A). Further analysis revealed that most types of adaptive immune cells (activated/central memory/effector memory CD4+/CD8+T cells, Tgd cells, Th1 cells, Treg cells, follicular helper T cells (Tfh), and activated/immature/ memory B cells) and cell types related to innate immunity (neutrophils, monocytes, macrophages, mast cells, activated, immature dendritic cells (DCs), NK cells, natural killer T cells (NKT), and MDSCs) are positively correlated with PPIB expression levels in LGG (Figure 4B). These results imply that NUTF2 promotes tumor progression by regulating infiltrating immune cells in LGG.
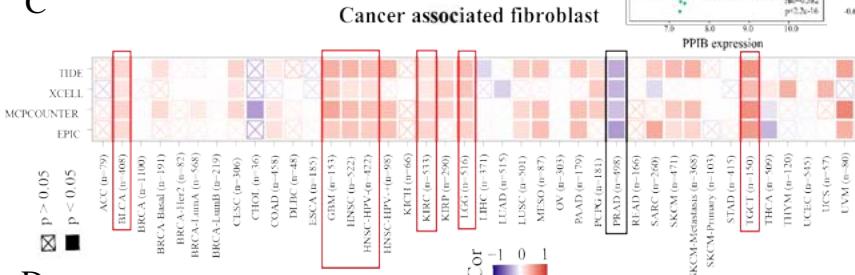
A kind of highly heterogeneous and hyper-activated fibroblast named CAFs has been demonstrated to promote tumor initiation, migration, inflammation, and drug resistance via the secretion of chemokines and cytokines, such as VEGFA and CXCL12(Guo et al., 2008 Jul 11; Nurmik et al., 2020 Feb 15). Our study utilized four algorithms (TIDE, XCELL, MCPCOUNTER, EPIC) based on the TIMER 2.0 tool to analyze the correlation between PPIB expression and CAFs in various tumors. The results revealed that the expression level of PPIB is positively correlated with CAFs in BLCA, GBM, HNSC, HNSC-HPV-, KIRC, LGG, and TGCT (appeared in 4 algorithms). By contrast, the
expression level of PPIB is negatively correlated with CAFs in PRAD (Figure 4C). These results implied that the correlation between PPIB expression and CAFs varies among different types of tumors.
 Panel label: A.
 Panel label: B.
 Panel label: C D.
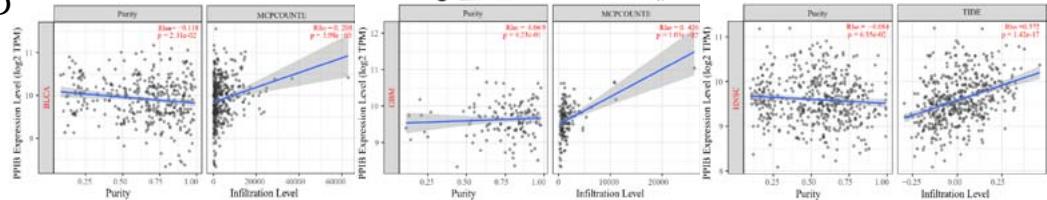
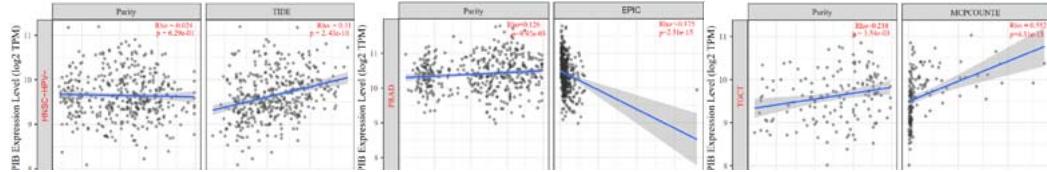 Figure 4: Analysis of PPIB in immune microenvironment.
(A) Correlation analysis between tumor infiltrating lymphocyte abundance and PPIB expression in tumors. (B) The expression of PPIB in LGG is correlated with the activation of CD8+T cells, central memory CD8+T cells, Tgd cells, CD56 positive cells, NKT cells, and monocytes. (C) Correlation between PPIB and CAFs calculated by four algorithms. (D) The top 6 tumors were positively correlated with expression level of PPIB.
### e) Enrichment Analysis of PPIB co-expression genes
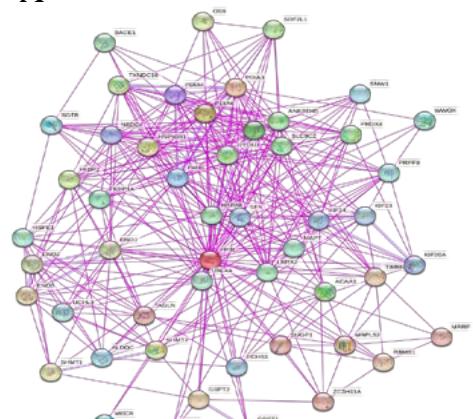
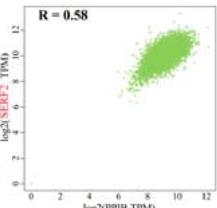
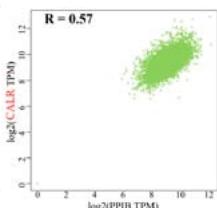
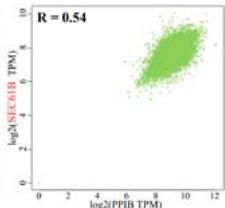
To investigate the potential molecular mechanisms of PPIB in tumor occurrence and development. In this study, we conducted protein-protein interaction network analysis (PPI) using the string online tool. As shown in Figure 5A, we obtained 50 PPIB binding proteins (Figure 5A). In addition, by combining the expression data of all TCGA tumors, we identified the top 100 genes most related to PPIB expression. Among them, five genes with high correlation with PPIB were screened: PDIA3 (R=0.64), MANF (R=0.63), SERF2 (R=0.58), CALR (R=0.57), and SEC61B (R=0.54). Moreover, we substituted the top 5 genes with high correlation into the TIMER2.0 database and analyzed that PPIB is positively correlated with their expression levels (Figure 5B).
We identified KEGG pathways related to cancer through functional enrichment analysis of the top 100 genes, such as protein processing in the endoplasmic reticulum, biosynthesis of N-glycans, and protein output
(Figure 5C). In addition, we conducted a Gene Ontology (GO) analysis on the relationship between PPIB and biological processes, cellular components, and molecular functions. Research has found that protein folding and protein disulfide isomerase activity may be involved in the role of PPIB in the cancer pathogenesis. These results revealed the possible molecular mechanisms and roles of PPIB in tumor pathogenesis (Figure 5D).
 Panel label: A C.
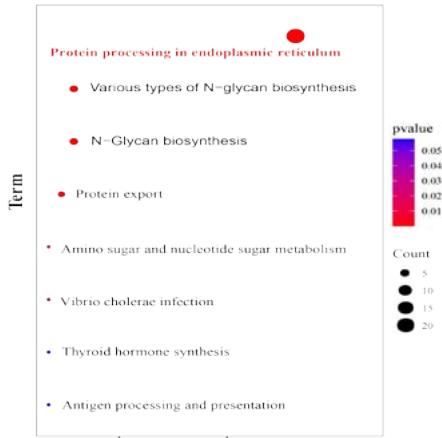 Panel label: KEGG PATHWAY.
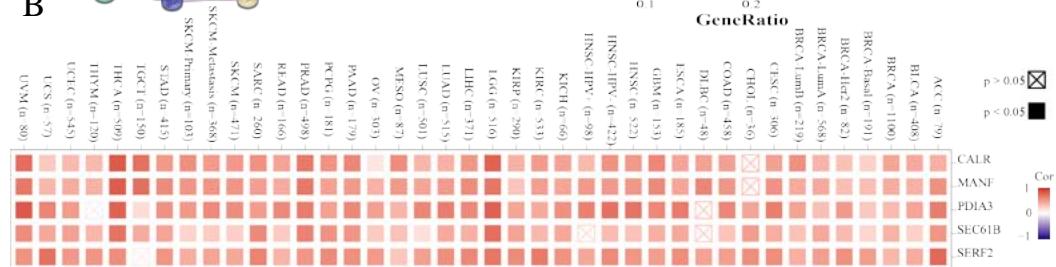 Panel label: B.
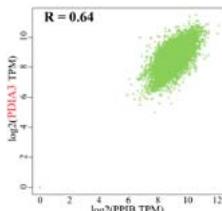
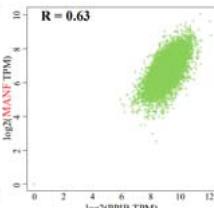
 Panel label: D.
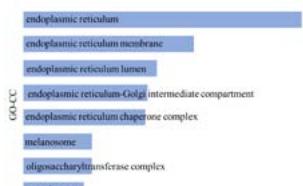

 Figure 5: Analysis of molecular mechanism of PPIB in tumorigenesis.
(A) Analysis of protein-protein interaction network of PPIB. (B) Correlation between PPIB and the first five genes in different types of tumors. (C) KEGG pathway analysis of the top 200 genes that associated with PPIB expression. (D) Gene ontology analysis of the top 200 genes.
## IV. DISCUSSION
Cancer has become one of the leading causes of increased human mortality. The process of cancer transformation is the result of cellular dysfunction. This is due to the accumulation of many genetic and epigenetic changes within cells, manifested as the accumulation of chromosomal or molecular abnormalities, leading to genetic instability (Peters & Gonzalez, 2018 Oct 1). For example, DNA methylation of SDC2 promotes tumor progression in colorectal cancer (Galamb et al., 2016).
Exogenous and endogenous factors, as well as individual factors, including genetic susceptibility, contribute to the occurrence and development of cancer. Environmental chemicals interact with chemical metabolism and endogenous signaling. Exogenous chemicals can form reactive intermediates with DNA through toxic metabolic enzymes. If not repaired by DNA repair enzymes, it can cause mutations in critical genes, such as tumor suppressors or oncogenes, ultimately leading to the formation of precancerous cells (Lewandowska et al.). Amplification is mechanism of activation of oncogenes such as fibroblast growth factor receptor 1 (FGFR1) and discoidin domain receptor 2 (DDR2) in SCC. Intriguingly, many of these genetic alternations are associated with smoking status (Cooper et al., 2013). The intrinsic molecular mechanisms of tumors are complex, and there is still a need to explore more potential molecular markers to study the mechanisms of tumor occurrence further.
This study conducted a multi-omics study on PPIB through pan-cancer analysis and found that, based on the TCGA and GTEx databases, the expression level of PPIB was significantly upregulated in 20 types of cancer compared to normal tissues. Moreover, high expression levels of PPIB are associated with poor prognosis. There are differences in the expression levels of PPIB among different pathological stages of KICH, KIRC, UCS, TGCT, and THCA. According to the above research, PPIB has a cancer-promoting effect. The high expression of PPIB in tumors indicates its potential as a tumor marker.
In the genetic variation study of PPIB, the expression level of PPIB is positively correlated with copy number amplification. But there is no significant correlation between PPIB mutations and prognosis. Therefore, we focused on the epigenetic mechanism analysis of PPIB. Methylation disorders are involved in many diseases, including human cancer. DNA methylation regulates the expression of target genes during transcription. Loss or silencing of promoter methylation leads to tumor progression (Dai et al., 2021 Mar 31; Ehrlich, 2019 Dec). Compared with normal tissues, the methylation level of PPIB is lower in multiple tumor types, and the expression level of PPIB is negatively correlated with the methylation level. Therefore, it can be considered that methylation of PPIB may be one of the important epigenetic mechanisms in tumor progression.
With the emergence and development of tumors, a series of soluble factors promote the influx of non-malignant cells (immune cells), blood vessels, and stroma, collectively becoming the tumor immune microenvironment (TME) (Bilotta et al., 2022 Oct 20). According to Weinberg et al, among the top ten characteristics of tumors, promoting angiogenesis, activating infiltration and metastasis, avoiding immune attacks, and promoting tumor inflammation are correlated with TME. These become essential factors in cancer cell behavior and disease progression (Hanahan & Weinberg, 2011 Mar 4). Immune cells and CAFs are essential components of TME. The importance of dynamically regulating cancer progression and influencing treatment outcomes is now widely recognized, and multiple treatment methods targeting various components of TME have been developed in recent years. Immunotherapy can inhibit the occurrence and development of tumors (Bejarano et al., 2021 Apr). The expression level of PPIB in most tumors was significantly correlated with most infiltrating immune cells in this study. Studies have shown that infiltration of CD8+T cells exerts anti-tumor effects, while B cell nuclear plasma cells can synergistically exert anti-tumor effects(Han et al., 2020 Jun; Wouters et al., 2018 Dec 15). Egelston et al. proved that CD8+tumor infiltrating lymphocytes are related to the excellent prognosis of triple-negative breast cancer (TNBC) (Egelston et al., 2022 Feb 8). CAFs are significant prognostic factors and therapeutic targets (Chen et al., 2021 Dec). In this study, the expression levels of PPIB were positively correlated with CAFs in BLCA, GBM, HNSC, HNSC-HPV-, KIRC, LGG, and TGCT. PPIB may be involved in the transformation and activation of CAFs. Contrary to the fact that PPIB plays anoncogenic role in most tumors, the results above indicate that PPIB in PRAD is negatively correlates with Treg cells and CAFs. It can be inferred that PPIB may play an anti-cancer role in PRAD. However, further specific research is needed.
This study explored the potential molecular mechanisms of PPIB's role in tumors through enrichment analysis of PPIB co-expressed genes. It was found that there was a significant correlation between PPIB and endoplasmic reticulum-related activities. Hasel et al. proposed that PPIB is located in the endoplasmic reticulum and is inhibited by cyclosporine A. Inhibition of PPIB in the endoplasmic reticulum may reduce the number of correctly folded proteins that ultimately reach the cell surface (Hasel KW et al., 1991 Jul). This research result is consistent with the PPIB mechanism research results. Wei X et al. found that cyclosporine A-mediated downregulation of PPIB, which can induce $\beta$ apoptosis of pancreatic islet cells and occurrence of endoplasmic reticulum stress (Wei et al., 2018 Nov). It can be inferred that PPIB has a potential role in the endoplasmic reticulum stress pathway, and its value as a targeted factor for promoting cancer mechanisms is highly worthy of further research.
This study is the first to explore the mechanism of PPIB in tumors through pan-cancer analysis, statistical analysis of differential expression of PPIB, clinical prognosis, genetic analysis, immune infiltration analysis, and enrichment analysis. PPIB is differentially expressed in most tumors and is associated with a poor prognosis. This study links its mechanism of action in the endoplasmic reticulum and infers its potential and value as a targeted factor in tumor treatment. Based on the high correlation between PPIB and endoplasmic reticulum stress pathway, we will further explore the specific location and mechanism of PPIB in the endoplasmic reticulum stress pathway.
### ACKNOWLEDGMENTS
We would like to thank the authors of previous studies and the staff members of the cBioPortal and
TCGA for providing available data. We thank the authors of previous studies for providing available data.
Authors' contributions
Study administration, validation, and design: Shuwei Ren, Yongsheng Huang and Haiyan Huang. Methodology, acquisition, and interpretation of data: Yan Ouyang, QiDai and Shengming Lai.
Writing-original manuscript: Yan Ouyang.
Study supervision: Shuwei Ren.
All authors read and approved the final manuscript.
Funding
This study was supported by funding from the National Natural Science Foundation of China (No.82203703 to Shuwei Ren) and the Guangdong Basic and Applied Basic Research Foundation (2021A1515111138 to Yongsheng Huang).
Data availability
Data are available in a public, open access repository.
Declarations
Ethics approval and informed consent
The study was conducted in accordance with the Helsinki Declaration.
Consent for publication
Not applicable.
Competing interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Availability of data and materials
All of the data in this study were described in the "Methods" section.
Generating HTML Viewer...
References
34 Cites in Article
Leire Bejarano,Marta Jordāo,Johanna Joyce (2021). Therapeutic Targeting of the Tumor Microenvironment.
Maria Bilotta,Antonella Antignani,David Fitzgerald (2022). Managing the TME to improve the efficacy of cancer therapy.
Freddie Bray,Mathieu Laversanne,Elisabete Weiderpass,Isabelle Soerjomataram (2021). The ever‐increasing importance of cancer as a leading cause of premature death worldwide.
Ethan Cerami,Jianjiong Gao,Ugur Dogrusoz,Benjamin Gross,Selcuk Sumer,Bülent Aksoy,Anders Jacobsen,Caitlin Byrne,Michael Heuer,Erik Larsson,Yevgeniy Antipin,Boris Reva,Arthur Goldberg,Chris Sander,Nikolaus Schultz (2012). The cBio Cancer Genomics Portal: An Open Platform for Exploring Multidimensional Cancer Genomics Data.
Darshan Chandrashekar,Bhuwan Bashel,Sai Balasubramanya,Chad Creighton,Israel Ponce-Rodriguez,Balabhadrapatruni Chakravarthi,Sooryanarayana Varambally (2017). UALCAN: A Portal for Facilitating Tumor Subgroup Gene Expression and Survival Analyses.
Yang Chen,Kathleen Mcandrews,Raghu Kalluri (2021). Clinical and therapeutic relevance of cancer-associated fibroblasts.
Tae Choi,Minh Nguyen,Jieun Kim,Yong Jo,Miran Jang,Ngoc Nguyen,Hyeong Yun,Wonchae Choe,Insug Kang,Joohun Ha,Dean Tang,Sung Kim (2018). Cyclophilin B induces chemoresistance by degrading wild‐type p53 via interaction with MDM2 in colorectal cancer.
W Cooper,S Lam Dc Fau -O'toole,Sa O'toole,J Fau -Minna,J Minna (2013). Molecular biology of lung cancer.
Xiaofeng Dai,Tiejun Ren,Yuxin Zhang,Nan Nan (2021). Methylation multiplicity and its clinical values in cancer.
G Dennis,Sherman Jr,D Bt Fau -Hosack,Hosack Da Fau -Yang,J Yang,J Fau -Gao,W Gao,W Fau -Lane,H,Lane Hc Fau -Lempicki,R Lempicki,R (2003). DAVID: Database for Annotation, Visualization, and Integrated Discovery.
Colt Egelston,Weihua Guo,Jiayi Tan,Christian Avalos,Diana Simons,Min Lim,Yinghui Huang,Michael Nelson,Arnab Chowdhury,Daniel Schmolze,John Yim,Laura Kruper,Laleh Melstrom,Kim Margolin,Joanne Mortimer,Yuan Yuan,James Waisman,Peter Lee (2022). Tumor-infiltrating exhausted CD8+ T cells dictate reduced survival in premenopausal estrogen receptor–positive breast cancer.
Melanie Ehrlich (2019). DNA hypermethylation in disease: mechanisms and clinical relevance.
Orsolya Galamb,Alexandra Kalmár,Barbara Barták,Árpád Patai,Katalin Leiszter,Bálint Péterfia,Barnabás Wichmann,Gábor Valcz,Gábor Veres,Zsolt Tulassay,Béla Molnár (2016). Aging related methylation influences the gene expression of key control genes in colorectal cancer and adenoma.
Xiaoying Guo,Hiroko Oshima,Takanori Kitmura,Makoto Taketo,Masanobu Oshima (2008). Stromal Fibroblasts Activated by Tumor Cells Promote Angiogenesis in Mouse Gastric Cancer.
Jichang Han,Nikhil Khatwani,Tyler Searles,Mary Turk,Christina Angeles (2020). Memory CD8+ T cell responses to cancer.
Douglas Hanahan,Robert Weinberg (2011). Hallmarks of Cancer: The Next Generation.
Karl Hasel,James Glass,Martin Godbout,J Sutcliffe (1991). An Endoplasmic Reticulum-Specific Cyclophilin.
Marta Kulis,Manel Esteller (2010). DNA Methylation and Cancer.
Anna Lewandowska,Marcin Rudzki,Sławomir Rudzki,Tomasz Lewandowski,Barbara Laskowska (2019). Environmental risk factors for cancer – review paper.
Ting Li,Hanqing Guo,Xiaodi Zhao,Jiang Jin,Lifeng Zhang,Hong Li,Yuanyuan Lu,Yongzhan Nie,Kaichun Wu,Yongquan Shi,Daiming Fan (2017). Gastric Cancer Cell Proliferation and Survival Is Enabled by a Cyclophilin B/STAT3/miR-520d-5p Signaling Feedback Loop.
Taiwen Li,Jingxin Fu,Zexian Zeng,David Cohen,Jing Li,Qianming Chen,Bo Li,X Liu (2020). TIMER2.0 for analysis of tumor-infiltrating immune cells.
Hidewaki Nakagawa,Masashi Fujita (2018). Whole genome sequencing analysis for cancer genomics and precision medicine.
Martin Nurmik,Pit Ullmann,Fabien Rodriguez,Serge Haan,Elisabeth Letellier (2020). In search of definitions: Cancer‐associated fibroblasts and their markers.
Xin Pan,Xiaoxin Ma (2020). A Novel Six-Gene Signature for Prognosis Prediction in Ovarian Cancer.
L Peddada,J Mcpherson,R Law,J Wasmuth,P Youderian,R Deans (1992). Somatic cell mapping of the human cyclophilin B gene (PPIB) to chromosome 15.
Jeffrey Peters,Frank Gonzalez (2018). The Evolution of Carcinogenesis.
Beibei Ru,Ching Wong,Yin Tong,Jia Zhong,Sophia Zhong,Wai Wu,Ka Chu,Choi Wong,Chit Lau,Ian Chen,Nam Chan,Jiangwen Zhang (2019). TISIDB: an integrated repository portal for tumor–immune system interactions.
R Stratton,M Campbell,J,P Futreal,P Andrew (2009). The cancer genome.
Hyuna Sung,Jacques Ferlay,Rebecca Siegel,Mathieu Laversanne,Isabelle Soerjomataram,Ahmedin Jemal,Freddie Bray (2021). Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries.
Zefang Tang,Boxi Kang,Chenwei Li,Tianxiang Chen,Zemin Zhang (2019). GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis.
C Von Mering,M Huynen,D Fau -Jaeggi,D Jaeggi,S Fau -Schmidt,P Schmidt S Fau -Bork,P Bork,B Fau -Snel,B Snel (2003). STRING: a database of predicted functional associations between proteins.
Xiao Wei,Dan Zhu,Chenchen Feng,Guofang Chen,Xiaodong Mao,Qifeng Wang,Jie Wang,Chao Liu (2018). Inhibition of peptidyl‑prolyl cis‑trans isomerase B mediates cyclosporin A‑induced apoptosis of islet β cells.
Wouters,M Auid-Orcid,& Nelson,B Liu,C,-O Hu,F,-O Xie,G,-O Miao,Y,-O Li,X Zeng,Y Guo,A,-O (2018). GSCA: an integrated platform for gene set cancer analysis at genomic, pharmacogenomic and immunogenomic levels.
No ethics committee approval was required for this article type.
Data Availability
Not applicable for this article.
How to Cite This Article
Yan Ouyang. 2026. \u201cPPIB Unveiled: A Comprehensive Pan-Cancer Exploration Unraveling Immunological Signatures and Prognostic Implications\u201d. Global Journal of Medical Research - F: Diseases GJMR-F Volume 24 (GJMR Volume 24 Issue F1).
Explore published articles in an immersive Augmented Reality environment. Our platform converts research papers into interactive 3D books, allowing readers to view and interact with content using AR and VR compatible devices.
Your published article is automatically converted into a realistic 3D book. Flip through pages and read research papers in a more engaging and interactive format.
Our website is actively being updated, and changes may occur frequently. Please clear your browser cache if needed. For feedback or error reporting, please email [email protected]
Thank you for connecting with us. We will respond to you shortly.